Output data¶
QRS - Internal in Igor¶
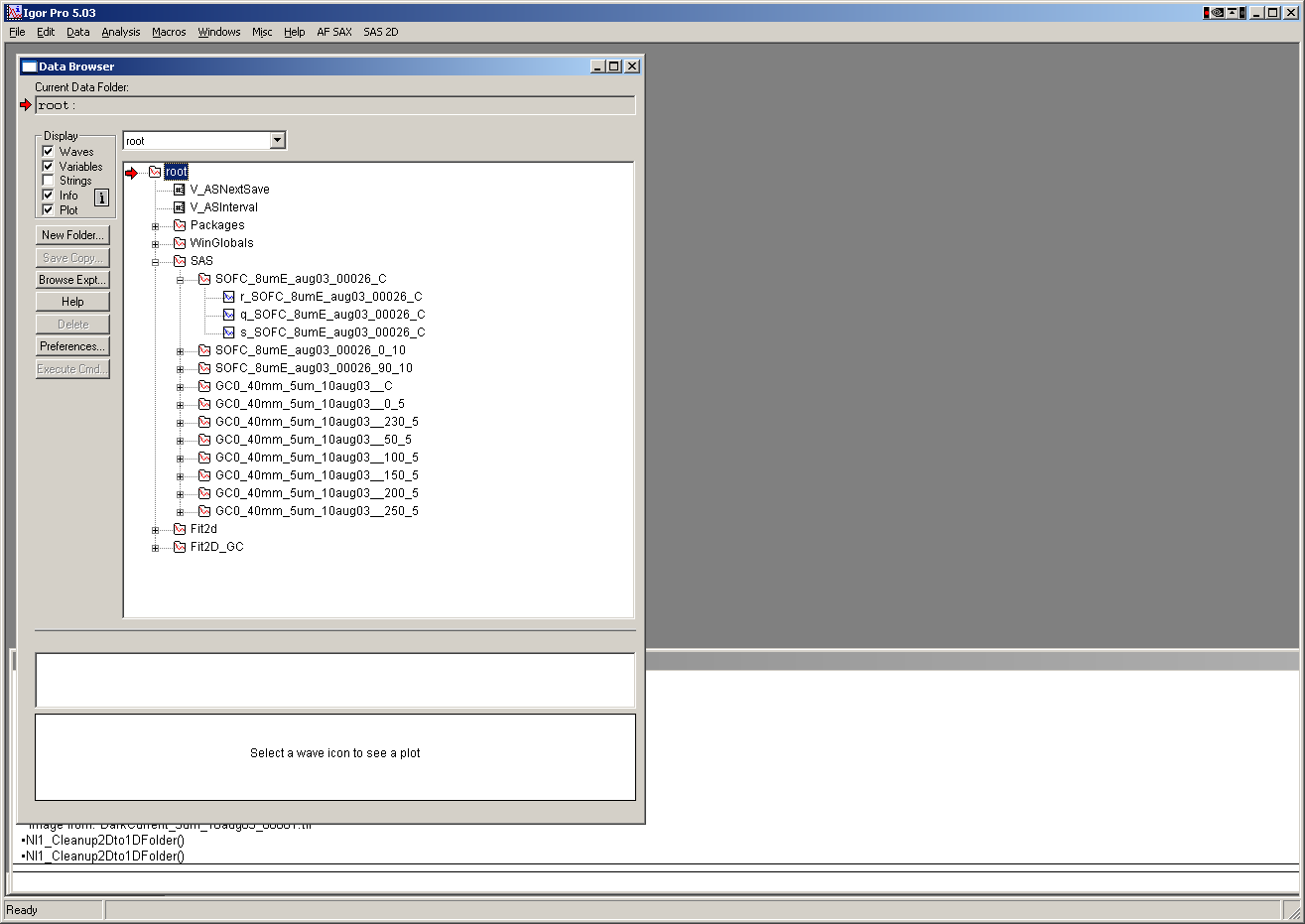
Data are internally stored (if selected) within Igor experiment in folder root:SAS: in folders with
nameOfSample_C being the circular average
nameOfSample_Angle_halfWidth being sector average around direction Angle with sector half-width.
The wave names
X-axis data
q_NameOfSample_C (or _Angle_halfWidth) q vector in A:sup:`-1`
t_ NameOfSample_C (or _Angle_halfWidth) 2 theta, if output with respect to 2 theta
d_ NameOfSample_C (or _Angle_halfWidth) d for output wrt d
y axis data
r_ NameOfSample_C (or _Angle_halfWidth) intensity (if calibrated in whatever units – thickness is converted to cm, so it should be cm:sup:`-1`)
error
s_ NameOfSample_C (or _Angle_halfWidth) error for intensity
other
w_ NameOfSample_C (or _Angle_halfWidth) width of each bin of Q/d.2 theta. This is for LUT output, and provides data for bin-width smearing. Smaller number of bins, larger width of each. For linear binning, this is same number and is (Max-Min/numOfPoits), but for log binning this is varying function of bin position.
For Line profile data:
For example for GI_Vertical line in my test case, this was the name:
gc_saxs_395__GI_VLp_0.0077
“gc_saxs_395_”…. Part of the name of used image
GI_VLp_.... GI_Vertical Line
0.0077 …. q:sub:`y` value at which the data were calculated.
Exported data are Int, error, Q, qx, qy, qz columns with header and column names
Saved data in Igor are
r_NameOfSample_ProfileIndicator_Qvalue intensity
q_NameOfSample_ProfileIndicator_Qvalue q [A:sup:-1]
s_NameOfSample_ProfileIndicator_Qvalue error
qy_NameOfSample_ProfileIndicator_Qvalue qy [A:sup:-1]
qz_NameOfSample_ProfileIndicator_Qvalue qz [A:sup:-1]
qx_NameOfSample_ProfileIndicator_Qvalue qx [A:sup:-1] (generated ONLY if GI… profile is used)
Note, intensity wave has attached wave note, containing some useful information:
CalibrationFormula=1*((Sa2D));CurrentMaskFileName=A mask_mask;QvectorNumberPoints=300;CircularAverage=1;
ASCII export¶
ASCII files with following data are stored in the selected folder:
# CalibrationFormula=1*((Sa2D))
# CurrentMaskFileName=A mask_mask
# QvectorNumberPoints=300
# AngularSector=150
# AngularHalfWidth=5
0.01601654 0 0
0.0163735 1537 39.20459
0.01655496 1467 38.30144
0.01673842 1416 999.0073
0.01692392 1505 38.79433
The columns contain first q, second intensity and third error…