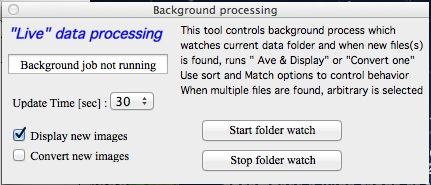
Main Panel¶
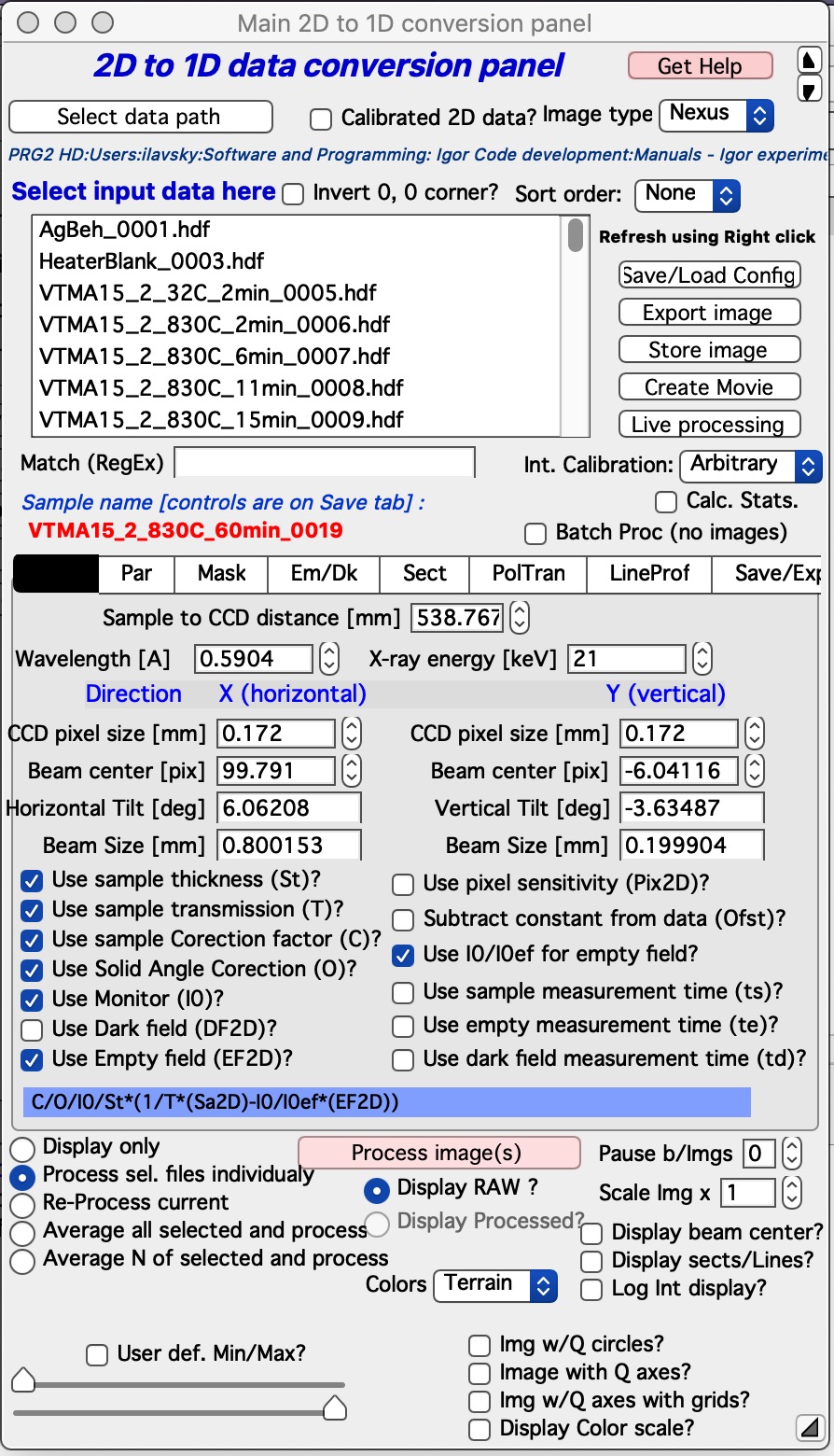
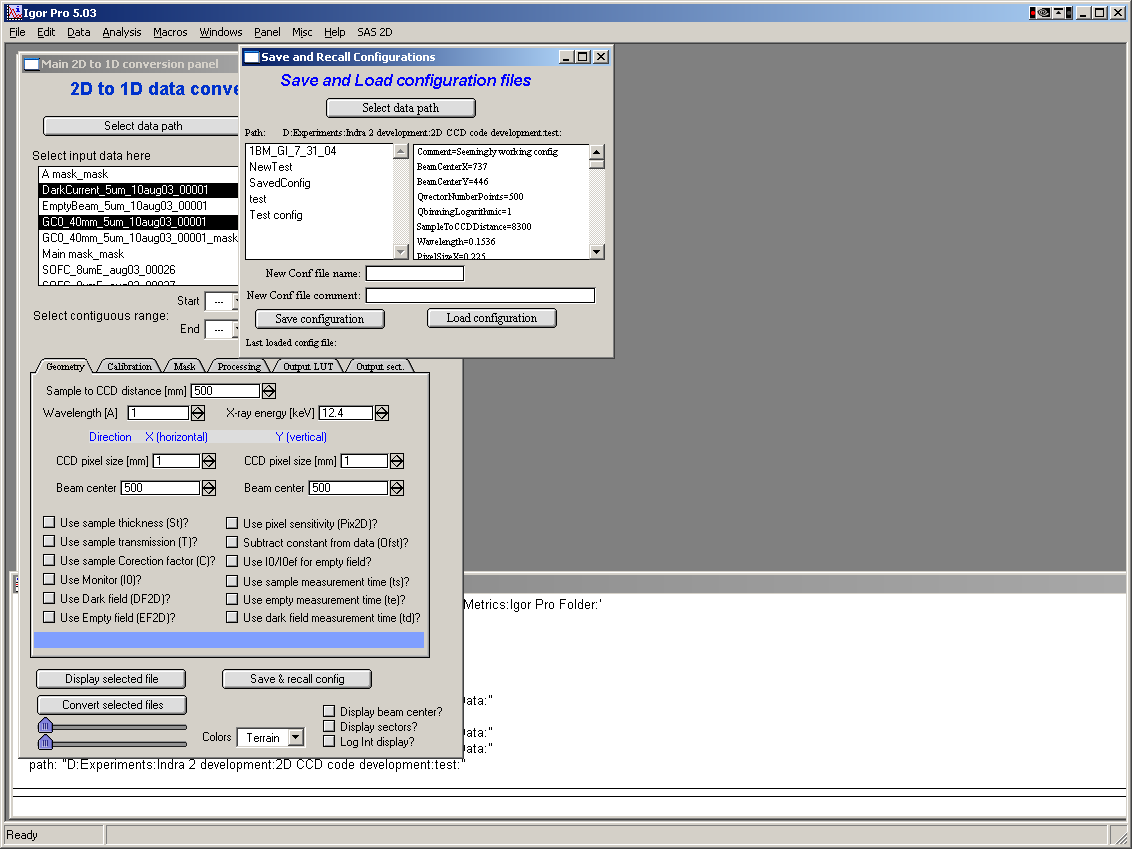
Select “Main panel” from the “SAS 2D” menu. This will present the following panel:
The panel has three major parts:
Top is designed for 2D data selection. Here user selects which 2D image will be processed.
Middle (tabbed area) is designed for controls of processing. This is the busiest area of the panel and each tab will be explained later.
Bottom contains buttons for main controls and 2D image controls.
Selecting data¶
Nika can load number of different image types - aka: file formats, file types - usually well described by file extension (e.g., tif). These are selected by “image type” popup menu in top right corner. If appropriate file type is not found in the “image type” popup menu, you will have to contact me so I can develop and add appropriate loader for your specific data. Note, that most data formats are binary data with some header, and if you can get description of your data format you can often use General Binary reader.
Select appropriate type of data you have and then push “Select data path” button, dialog is presented, in which path to folder on the hard drive containing 2D images is selected. Find the local path to data using this standard Igor dialog. and push OK when done.
NOTE the “Calibrated 2D data?” checkbox. If selected, Nika expects 2D calibrated data – fully normalized and corrected data provided as one of the 2D formats, basically 2D image of Intensity, Q (vector), and uncertainty. Number of options is being current developed, the code currently handles EQSAXS (ORNL) and canSAS/Nexus. This part is under heavy development at this time, expect changes…
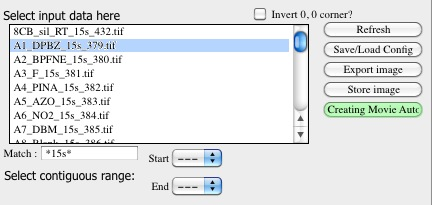
When valid path is selected, the Igor will check the folder and list all files of appropriate type (assuming the files have extensions) in the ListBox below the button.
Here user can select one files, more files (by holding down shift key on Windows) and continuing selection (using the two pull down menus below the list Box)…
Note, that from Nika ver. 1.66 Listboxes have right click actions and users can refresh content and perform some functions from right click.
Use the “Match” field to mask the file names with Regular expression. To match part of the name, just use the string needed - so matching samples with _15s in name, just add _15s in the field. Regular expressions are very powerful, read on line how to use them.
Note, the files ending with “_mask”. These are mask files created by Nika package, these were used to be tiff files, now they are hdf5 files… Separate chapter explains how mask is created.

Invert 0,0 corner¶
As default Igor displays 0,0 of the image in the top left corner. This seems to be distressing for some users, so if checked, images will have 0,0 in the left bottom corner. Nothing else is changed, so the orientation of sectors WRT original image is preserved and reduced data are the same as without this checkbox checked. Simply, the processing of Nika package is independent of this checkbox, it is ONLY cosmetic…
Sort order¶
Decides how the data are listed in the listbox. Options:
None – list as provided by OS.
Sort – the old method. Alphabetical (but numerical order may get wrong)
Sort2 – alphabetical, but taking care of sorting out smaller number before larger ones.
_001. – this one assumes, that end of file name, before extension, is number. Before number you need to have “_” and after number must be “.” followed by extension.
Invert _001
Invert Sort
Invert Sort2
All inverted sorting simply reverses the sorting logic. Try them and see, which works best for you.
Match¶
Using RegEx now. This is Grep language using regular expressions, very powerful. For simplicity: match names containing (anywhere) test, just type in this field test. To match names starting with test type in ^test. Names ending with tif can be matched by tif$ and so on. Note that to match any single character you need to use. Need to start quickly? See here: https://www.cheatography.com/davechild/cheat-sheets/regular-expressions/
Batch processing (no images)¶
This is way to significantly speed up processing of images in Nika.
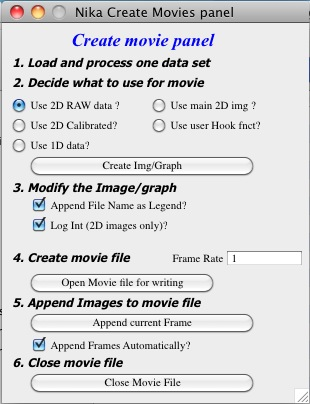
Testing has shown that up to 75% of time to process data in Nika can be spent on displaying the images, drawing into the images, and graphing the 1D data. And printing notes in the history area. Most of the time this is acceptable and images help users to understand what is happening. However, when processing large number of images this can needlessly slow down processing. The checkbox Batch Proc. (no images) speeds up processing by preventing needless image display. If this checkbox is selected, Nika will stop all image displays, updates of opened graphs and to indicate it is working will just display a panel Nika is batch Processing data (see next image). While this panel is up, Nika is running, but the only thing changing user can see is the red Sample Name on the main panel. When the selected batch of samples (batch is selected in the Select input data listbox) this panel will disappear.
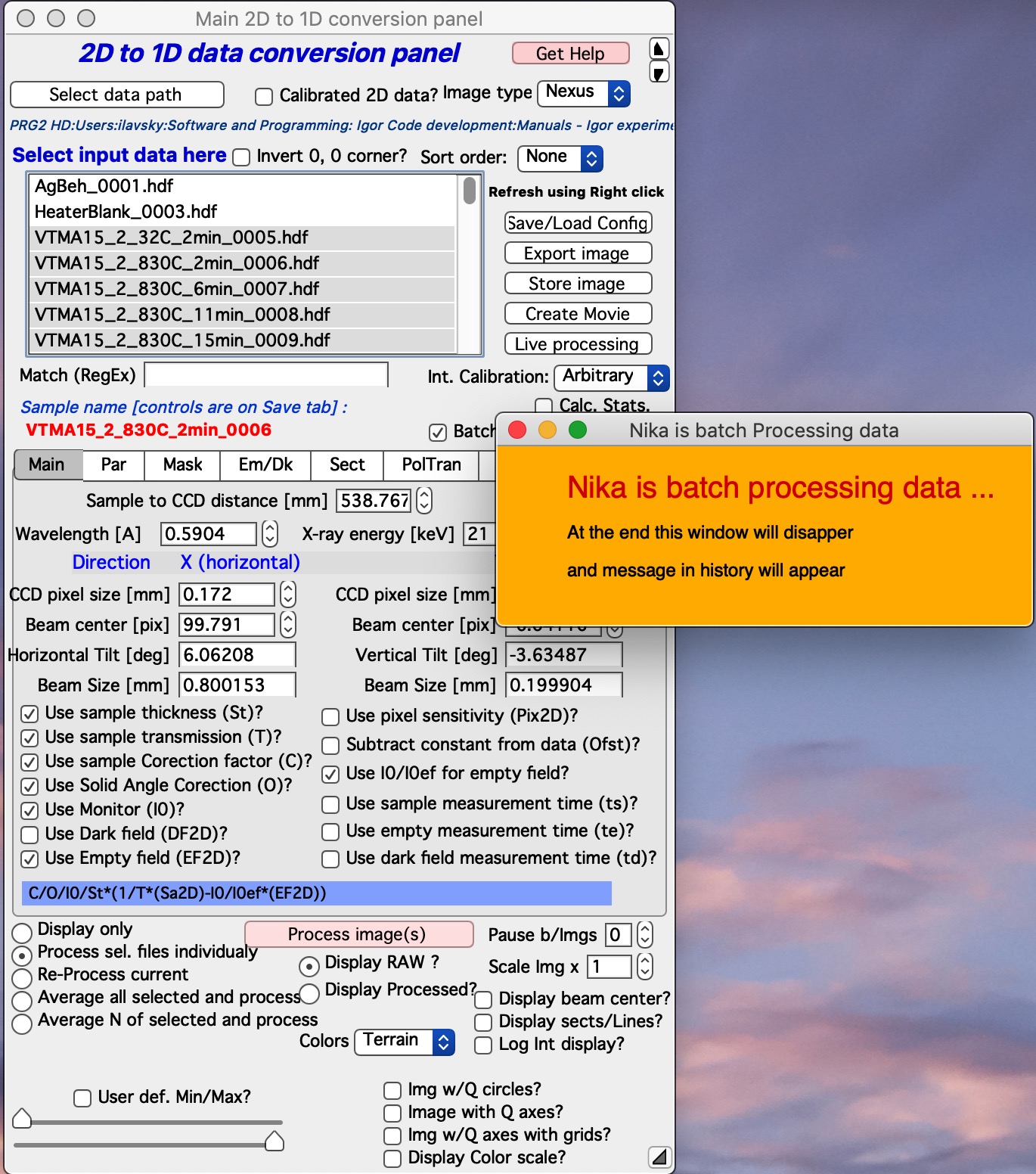
Also, notes are printed in the history area during the start and after the end of this batch processing :

Suggestion: Process one or two images first and verify the settings are correct and all parameters are correctly set. When you check the parameters and understand, that all is working right, you can run larger number of images in batch mode.
If the batch processing hits error and stops : Nothing bad happened. Close manually the panel Nika is batch Processing data (it can be killed as any other panel), fix the problem, and start again from where Nika stopped.
Controls in tabs¶
Note, that if images are averaged, they are first averaged during loading, and then – during processing to create lineouts / square matrix are corrected as described below. Therefore all parameters here related to single (if possibly averaged) image!
These are controls in the tabbed area.
We will now go through each tab separately
Main¶
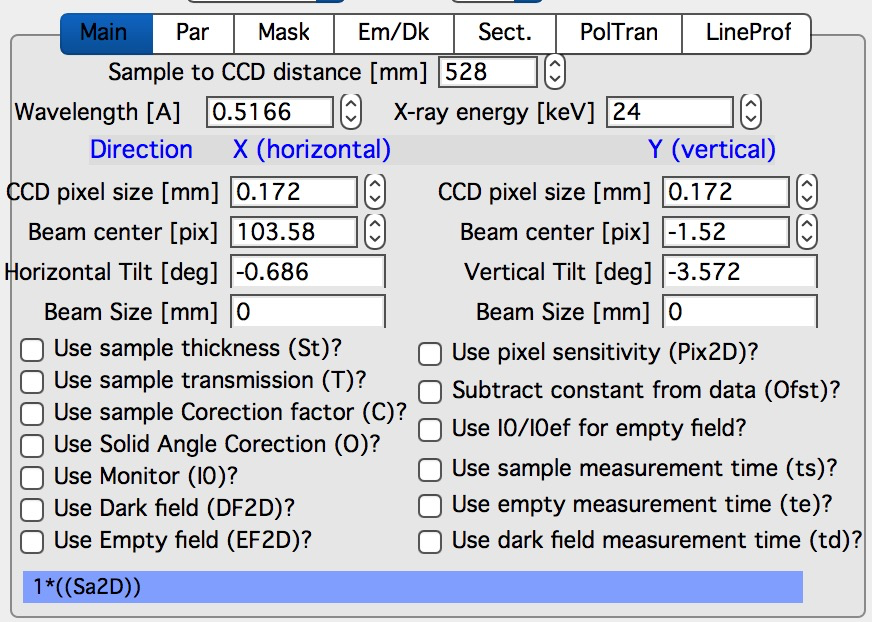
Here are some very clear parameters, related to SAXS camera geometry:
Sample to CCD distance in millimeters, Wavelength/Xray energy (these windows are linked), CCD image pixel size in mm (in X and Y directions). Note, X direction is horizontally, Y direction vertically. And Beam center position. Note, one can display beam center (to check it) in the graph by checkbox below the tab area.
And further there is pile of checkboxes, which describe method how to calibrate the data. Note, that formula used for calibration appears below to avoid any misunderstanding of the method. Select method needed for processing – and following tabs will have the appropriate controls available.
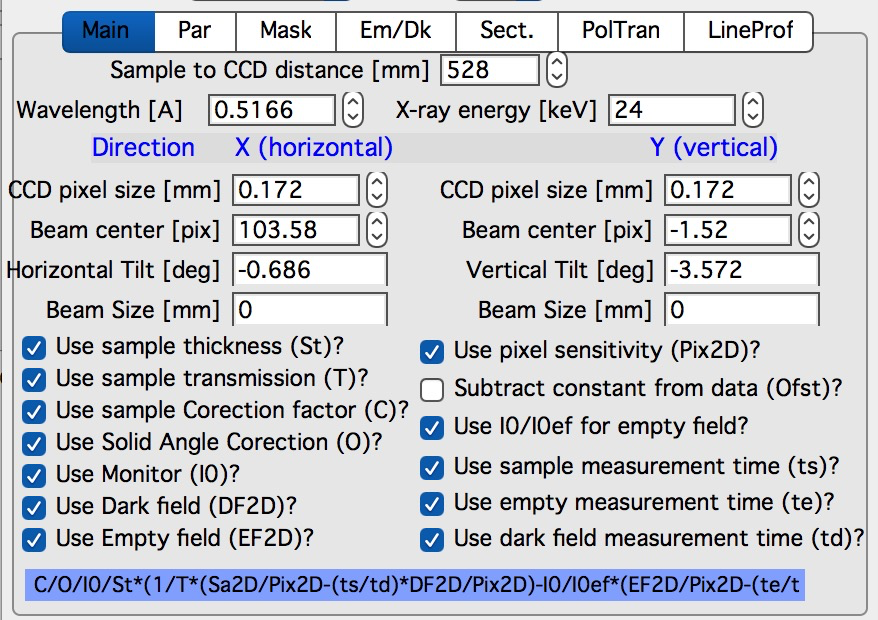
Note, that “Use of Dark field” and “Subtract constant from Data” cannot be used at the same time (they are effectively the same type correction)…
Note, only the appropriate controls will appear, so seeing all of these at the same time should be VERY unusual…
Comment for Use of Solid Angle Correction: When selected, the data are divided by solid angle of the central pixel (same value for all pixels). To correct for change in pixel solid angle as function of scattering angle, use Geometrical correction. Most of the time we do not bother with this option – if you use secondary calibration standard (like Glassy carbon or water) solid angle correction is included in the Calibration constant. If you do not use calibration and have relative data, you do not care also. The real need for this option is when you use data obtained in different sample to detector distances and want to combine the data together. Then this is necessary option.
Just remember, if you have obtained calibration constant, it is linked with the choice of the Solid angle correction.
Par¶
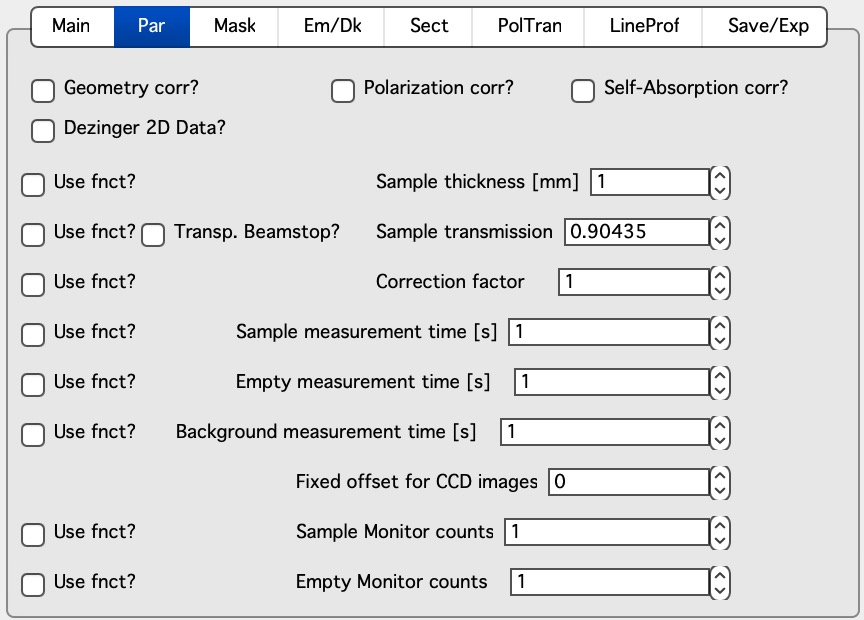
Here are standard controls (self explaining I hope):
“Geometry correction” – fixes the VARIATION of solid angle projection of the pixels on planar CCD detector. Mostly negligible for SAXS data… Just for completes, this divides the intensity at each pixel by (cos(2Theta))^3. And for those, who do not understand this formula, it took me may be 3 weeks to check it (I stole it from NIST data reduction). Very simplified, one cos(2theta) corrects for change of pixel radial direction as function of scattering angle, second cos(2theta) comes from change in distance between sample and detector as function of scattering angle in radial direction, third cos(2theta) comes from the same correction for tangential direction. Tangential size of pixel does not change as function of scattering angle.
“Polarization Correction” – Correction for either unpolarized radiation (desktop instruments with tube sources for example) or for Linearly polarized X-ray sources (synchrotrons). Opens up a new panel.
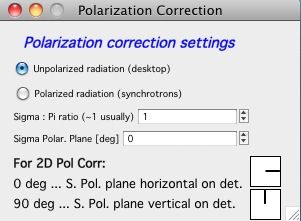
For unpolarized radiation use “Unpolarized radiation”. This is applicable ONLY to unpolarized radiation, the intensity data are corrected by formula:
Intensity_corrected = Intensity_measured / (0.5*(1+cos((2theta))^2))
For linearly polarized radiation use “Polarization radiation”, see separate chapter on Polarization correction little bit further in this manual.
By the way, for small-angle scattering each of these corrections is negligible.
“Dezinering” - Data, Empty, and Dark field images can be “dezingered” during loading. In this procedure each point is compared to surrounding pixels and if it is significantly larger (that is the dezinger ratio, if 2 then if the pixel is 2x larger than average of surrounding pixels) it is replaced with the average of the surrounding pixels. This is to remove spurious very high intensity points, which occur on some instruments.
It is possible to dezinger each image multiple times, in case the “zingers” are larger than single pixel.
Calibration/processing parameters:¶
Sample thickness in millimeters
*Important note: Nika versions prior 1.75 had a bug in the code, which caused the thickness to be used in mm and not converted into cm, as appropriate for SAXS data calibration. This was fixed in Nika version 1.75. BUT, this means, that calibration constants obtained on prior versions of Nika need to be also scaled by factor of 10 to account for this. I suggest carefully revising calibrations when upgrading to new version of Nika. This message will be also provided to users when new Nika version finds panel created by old Nika version. My apologies for this issue. Note: Under usual conditions when measurement of standard was reduced in Nika and then calibration constant was obtained this bug have cancelled out. This is also the reason why this bug was not found for so long. Thanks to a user, who actually read the code and found the bug.
Transmission as fraction. Note, if you have semi transparent beamstop, you can use Transp. Beamstop checkbox and input radius of the beamstop in pixels (read it from images with cursors). You need to have at least one sample image loaded in Nika. Nika will create a mask over current beam center and, when sample is being corrected, Nika will calculate average intensity in the circle of radius you provided around the beam center which was input at the time of checking this checkbox for sample and empty. Sample/Empty ratio is then transmission. Note, that if you change the beam center position, you need to rerun the code creating the mask for this calculation. Simply uncheck and check this checkbox “Transp. Beamstop” again.
Correction factor is for secondary calibration factor.
Measurements times in seconds, for each image.
Sometime one wants to use measurement time to correct images collected at different time exposures. While not suggested, it is possible to do here. I strongly discourage this.
Monitor counts allow scaling data by using monitor on incoming intensity.
“Fixed offset for CCD images” this is single value to be subtracted from each pixel of image to be processed.
*“Monitor counts”* use monitor counts to scale images (Sample/Empty)… This makes no sense for dark field…
Each of these values can be inserted by user as number, or using function:
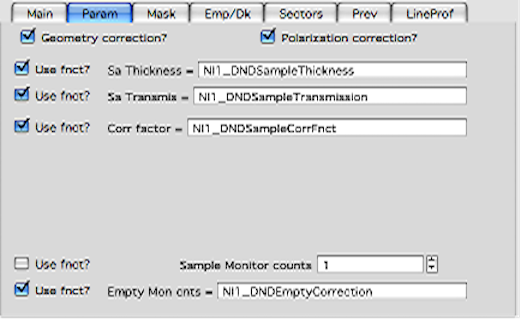
These function need to be “look up” functions, which are called with image name as parameter (FunctionName(“imageName”)) and must return single real number. The real use is to provide automatic look up of parameters from some records written by instrument. Above example is from included special support for DND CAT instrument.
Let me point out once more here, that using some of these corrections together makes no sense… Choose wisely.
Mask¶
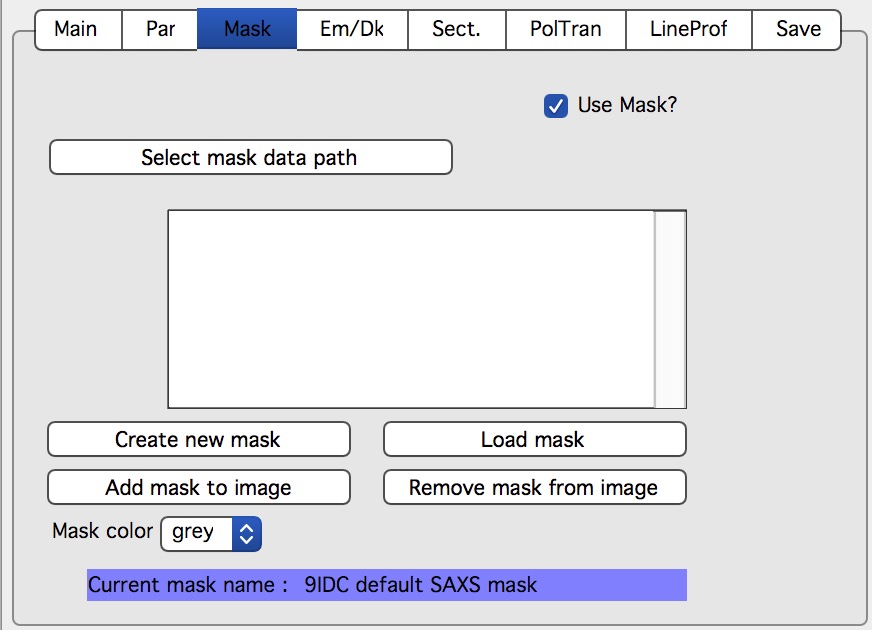
First checkbox, if Mask should be used (did not fit on the front tab…), button to select path to files with masks. Note, mask files created by Nika used to be always tiff files, with name in following manner: UserName_mask.tif Starting with version 1.49 they are now hdf5 files. These can be loaded in same as tiff files, but have anb advantage that these can be later modified in the mask tool…
Following are function of the buttons:
Create New mask – calls tool to create mask (see later in the manual)
Load mask – load file selected above in the list box as mask
Add mask to image – adds mask into the 2D image from the image
Remove mask from image – removes the mask from the image
Mask color – allows to change color (red, green, blue, black) of the displayed mask…
Current mask name – shows name of last loaded mask file
Emp/Dark¶
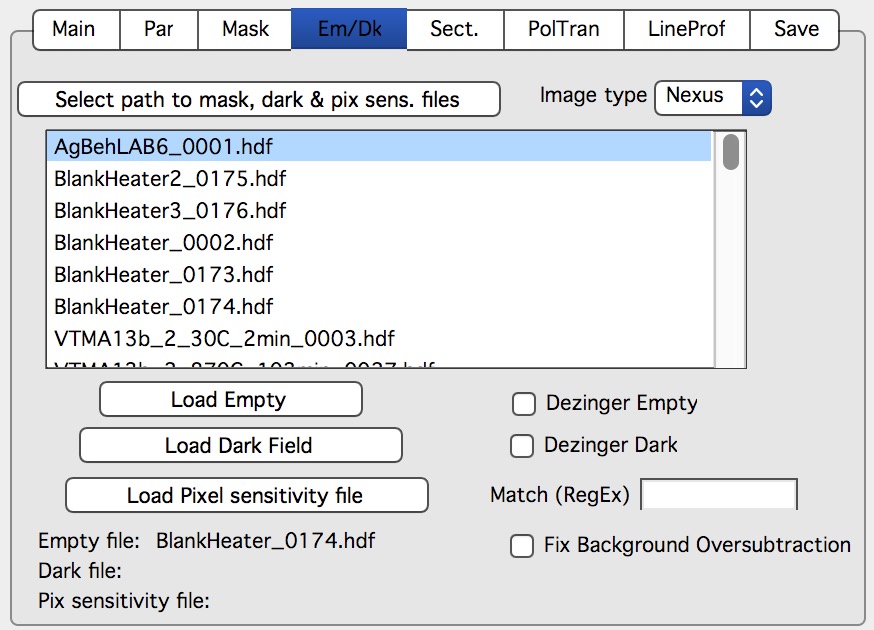
Here are controls for Empty/Dark field/pixel sensitivity (aka flood) images.
Button “Select path to mask, dark & pix sens, files” Selects path to data with the Empty, Dark field etc. I believe the files need to be the same type as data file (I need to check this).
Further buttons load the Empty/Dark/Pixel sensitivity, allow Dezingering of these (same method as the sample dezingering as selected above). And at the bottom are listed the file names of the files loaded…
“Fix Background Oversubtraction” - when checked, Nika will attempt to fix cases where background is stronger than sample+background scattering (after sample is fully processed). Another words, Nika will attempt to prevent negative intensities in processed data. This is done to prevent problems to downstream software, where some software (e.g. Irena) does not like negative intensity (which is physically meaningless). Now, this is bit tricky how to do this. This method is basically very simplistic, after processing, calibration, subtraction, etc., Nika will check if any point of resulting 1D data is negative. If so, Nika will find the most negative point (let’s ay it has intensity value of I1 < 0) and then add - to ALL points - 1.5 * abs(I1). This shifts the whole scattering intensity curve up by 1.5 * abs(I1), which may cause troubles for absolute intensity calibration etc if this value is not negligible compared to data with signal. So think hard if this is right. It works quite well for samples which have high-noise weak background at high-q.
Sectors¶

This tab controls how data are processed when method using “reverse Lookup tables” is used. This is the more suggested method for regular data processing. In this method Nika creates first lookup table for each sector defined and then can process much faster subsequent data files with the same geometry…
Controls:
Q space/d space/ 2 theta space – Output as function of Q, d, or 2 theta…
Min/Max (Q, d, 1 theta) range of evaluated Q, d, 2 theta. Set to 0 for automatic – automatic means, that the min/max is set for first q/d/2 theta which has non zero intensity
“Log binning” – check yes if Q/d/2 theta binning should be in logarithmic.
“Number of points” – number of points in Q/d/2 theta which should be created.
Do circular average – self-explanatory.
Make sector averages – do sector averages. Controls below control orientation and sizes of sectors. To see how the sectors are places, check the checkbox at the bottom of the control panel.
Create 1D graph – if checked, 1d graph with output is created (if necessary) and data added. Note, the graph may be crowded very fast, since data are added, and added…
Store data in Igor experiment – keep data (as qrs triplets) in current Igor experiment.
Overwrite existing data if exist – if data with the same name exist, overwrite without asking. Otherwise, you will be asked.
Export data – export ASCII data
Select output path – select where data are to be placed.
Use input data name for output – automatically name 1D data (with sector information added as DataName_Angle_width) by input data name.
ASCII data name – if the above is not selected, this is place to place name for output file. Note, if there is nothing available for the code as sample name, it will ask for some…
PolTrans¶
This means: ”Polar transformation” – prior (pre 1.68) name was “Preview” which is the intended use of this tool…
First:
This tool can use the calibrated data set (as well as RAW data set, depending on checkbox setting) so same calibration procedure is used as for the other processing. This tool is, however, less precise and does NOT produce useable errors. Be warned, this tool is meant as quick look on the data in different directions and not for final data processing…

This method is used to convert Intensity vs azimuthal angle from “polar coordinates” around beam center to plot where azimuthal angle is on vertical axis, pixel coordinate is on horizontal axis and intensity is expressed as color map. In here, one can produce rectangular graph:

On vertical axis is angle from 0 degrees axis (horizontally right from the beam center) and on horizontal axis is pixels distance from beam center. This is effectively set of lineouts in all azimuthal angles. It should be noted, that the code works very well for relatively small widths – may be up to 5 degrees, then the code becomes less precise, so keep angles small. Suggested is 1 -5 degrees.
These data then can be processed further by use of “image line profile” tool. This tool for now has it’s own “mindset” and does not properly update always. The dependencies are quite complex. If it does not update, close the tool and reopen…
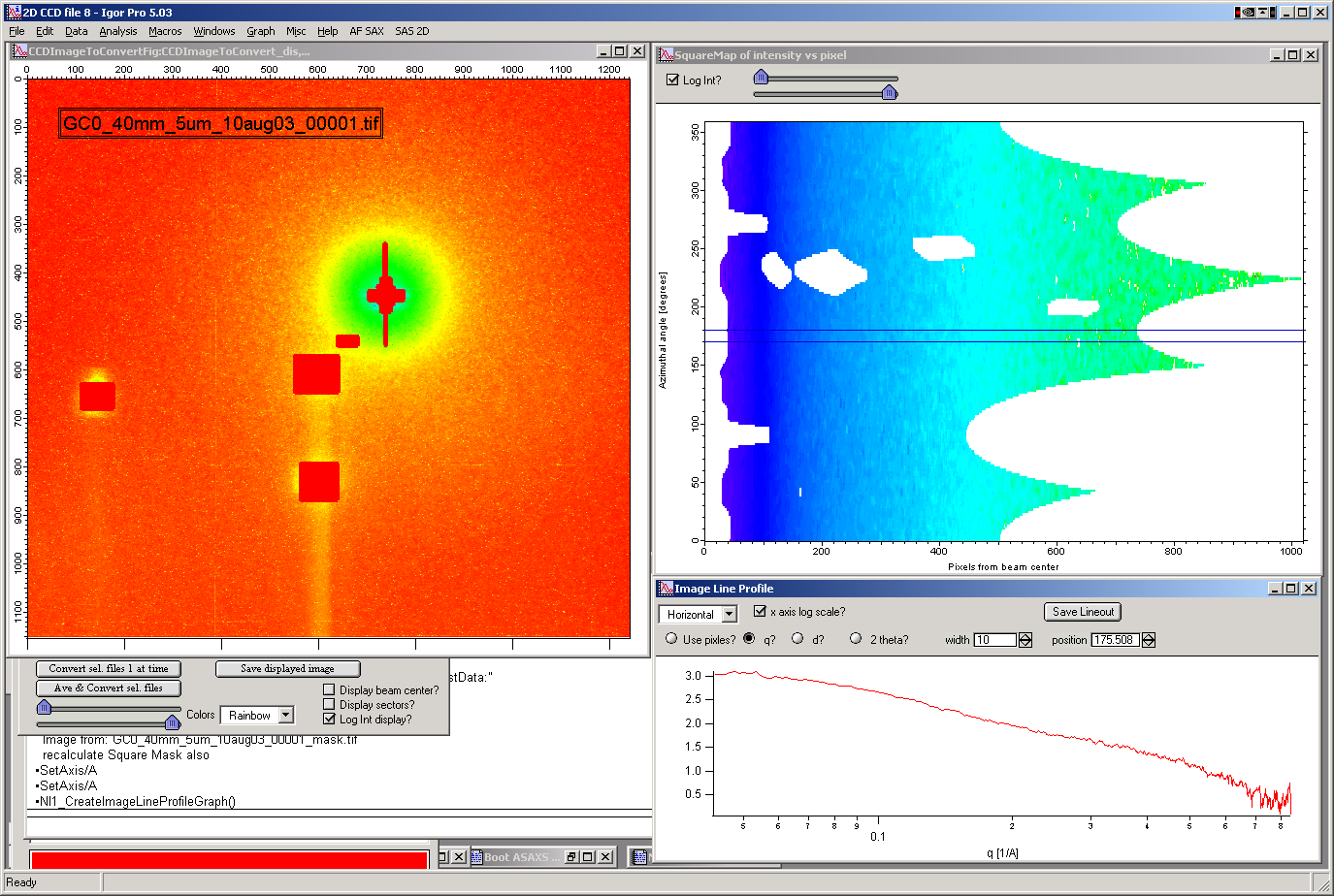
The “SquareMap of Intensity vs pixel” graph on the top right above shows the intensity in linear/log (checkbox left top corner) as function of pixel (bottom axis) and azimuthal angle (left axis). The lineout plot at the right bottom shows the intensity from this plot (note, the log/lin scaling in the image translates here!) as function of pixels/q/d/2 theta. Note, that this produces “natural” binning with every step in pixel is assigned single q/d/2theta position.
Note, the controls:
Number of sectors
Width of each sector - it is possible to have width such, that bins overlap, touch or do not touch… Default here is to have them touching.
Start Angle (0 = right horizontally from beam center)
End angle (wrt to start angle, most likely 360 degrees, or 180 degrees for only top half).
Mask data this tool does not mask, unless selected here…
Note, that by selecting larger width here, one can get very good and reliable sector average and manually move this average through the different azimuthal angles. Very useful, when hunting for particular azimuthal orientation…
Use RAW data if selected unprocessed image is used.
Use Processed data if selected processed image is used, available ONLY if the last image was loaded using one of the “Convert…” buttons, unavailable if the last image was loaded using “Ave & display sel. files(s). If the data were loaded using “Ave & display…” button, processed data do not exist.
Controls on Lineout tool:
Orientation of line profile (Horizontal/vertical)
X axis linear/log scale
Use: pixels/q/d/2 theta
Width and position
Save lineout – this saves “qrs” data in SAS folder in current Igor experiment. Suggested folder/data name is offered through dialog and user can modify as needed. Note, that errors are simple sqrt(intensity) – another words, these errors are not very useful.
LineProf¶
This tool calculates Intensity profile along curve on the detector. It uses different method than Sectors tool. Therefore, there are some important differences in how to use this tool…
The differences:
“Sectors” use inverse lookup method and can be set to create multiple different sectors on one image at once. Since this tool caches the lookup tables, it is slower first time, but much faster on subsequent images. This tool can be used ONLY by setting the data reduction parameters and then using buttons “Convert…”. You cannot manually evaluate any sector and no preview is provided. This tool causes high memory sizes of the Igor experiments with Nika package – the lookup tables are large. But it is fast for what it does.
And you can setup multiple sectors to be evaluated at once.
“LineProf” uses built in Igor Line Profile tool. It can be set ONLY to process one line profile at a time. This tool does not cache anything, so it takes the same time to process for each image. However, it is relative fast and can be used manually on Converted image. So, there are two methods to use it:
Set one line profile parameters, choose how to save data and push one of buttons “Convert..”
Do not set any conversion parameters, but use one of the buttons “Convert..”, set the LineProf tool to use Processed data and then set parameters for the
You can only set one line profile at a time, unless you manually create multiple profiles on each converted image.
Controls:
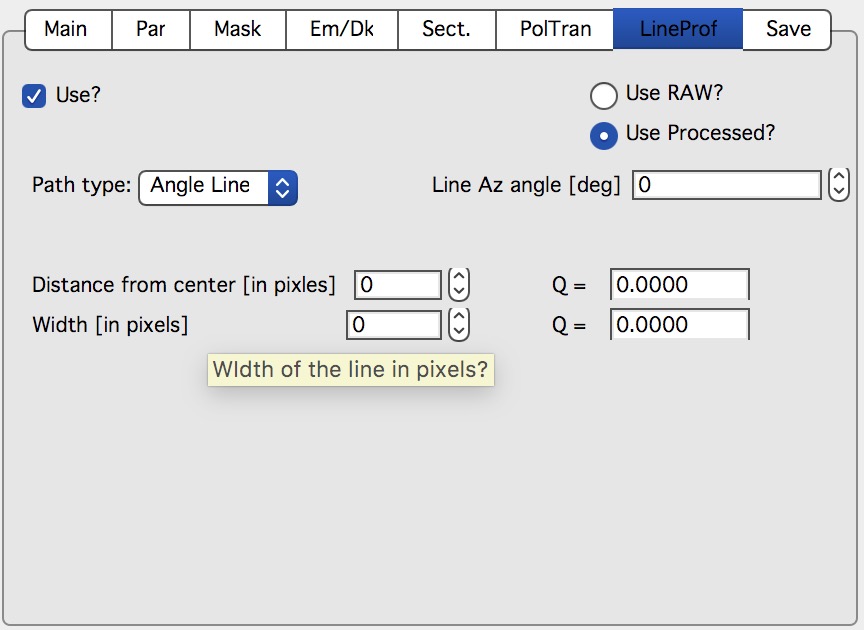
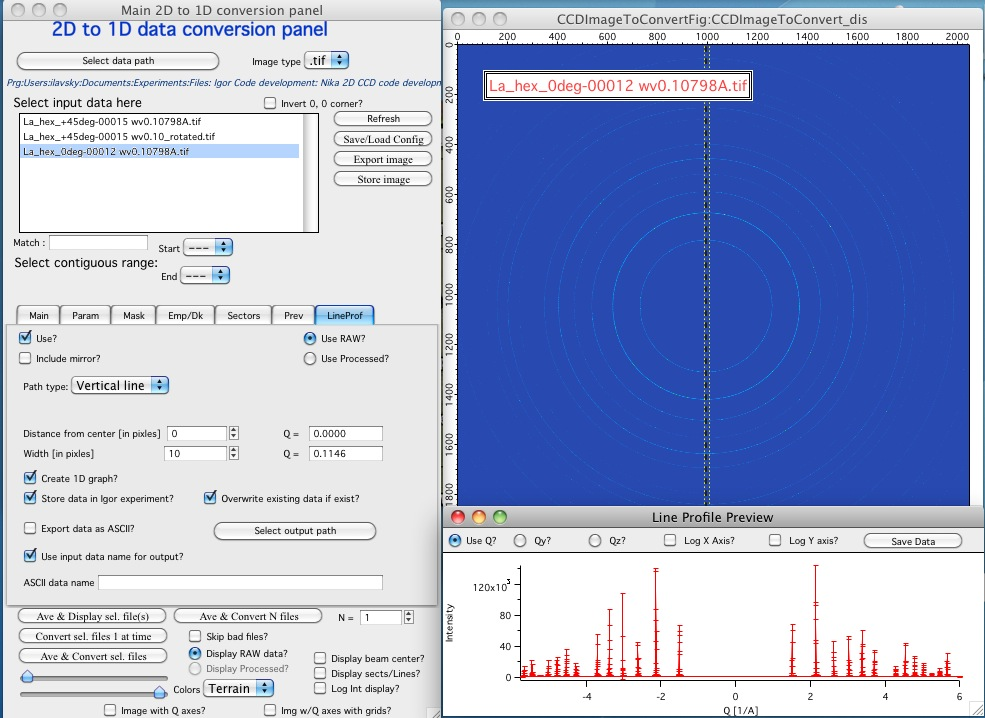
NOTE: some controls from the lower graph tab are moved to next tab, so this image is slightly obsolete. Will be fixed later.
New controls here:
“Use?” – switches on this tool.
“Use Raw?” – and “Use Processed?” – choices which image the tool will be used on. User Processed is not available if the last data set was loaded using “Ave & Display..” button (no Processed data are created in this case). NOTE: if you hit any button
“Convert..” and this tool is enabled, it is set to “Use Processed” automatically.
“Distance from Center [in pixels]” – user control to move the object to specific q . The q where the data will be calculated is displayed next to this control and is the appropriate q (q:sub:`y` or q:sub:`z`) for give shape. See Ellipse definition for specific there. NOTE: you must control the pixel position. Positive direction is to the right of the beam center (horizontally) or up from the beam center (vertically). Lines are drawn to help user image this out.
“Width [in pixels]” – width of the profile (minimum used one is 1 even if 0 is set by user) in pixels. This is the control to use to change how wide stripe is averaged. Next to it is control which shows this in q units. NOTE: the q width is calculated simply by subtracting Q values for the sides of the stripe. Intensity is averaged at each point perpendicularly to the direction of the line (curve). If more than 1 pixel is used for averaging, standard deviation of average is provided as error, if only 1 pixel is used, square root is used (which may be seriously WRONG)… You were warned.
This tool calculate intensity, intensity uncertainty and q, q:sub:`y`, and q:sub:`z` values. If one of GI profiles is used, it will calculate q, q:sub:`y`, q:sub:`z`, and q:sub:`x` values. See below.
IMPORTANT:
Of course, GISAXS community had to adopt different definition of Qx, Qy,a nd Qz than I did years ago, and therefore, this tool uses somehow different definitions than rest of Nika. So the horizontal direction (x-direction for Nika) is the Qy direction. Vertical direction on the detector is “y” direction for Nika; but it is direction of Qz. Please, keep this in mind… For those adventurous souls, who actually read my code, keep in mind at some point the code switches on your the x-y image coordinates to y-z-(x) GISAXS coordinates… Sorry. No other fix I would know about.
For now these are the available profiles:
*Vertical/Horizontal line:*
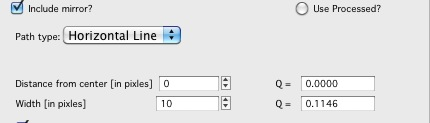
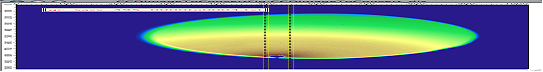
There is one more control available – “include mirror” (above the popup). If this is selected, mirror line over the beam center is included in calculations, see above.
This is line profile for transmission geometry.
Angle line:
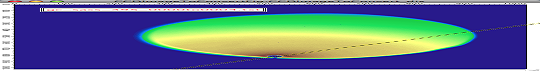


This is also for transmission geometry.
*GI_Vertical line & GI_Horizontal line*
These profiles are for Grazing incidence geometry. They need Grazing incidence angle:
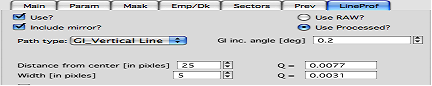
Both can include mirror image line across the beam center.
Note, that the position is defined in pixels as before, but the Q values are corrected according to the Grazing incidence geometry corrections, see Gilles Renaud, Remi Lazzari, and Frederic Leroy, Probing surface and interface morphology with GISAXS, Surface Science Reports 64(2009) 255-380, formula (1).
Note: before version 1.68 there was bug in the code for calculation of one of these angles. It hopefully had negligible impact for higher angles, but for small angles the Q calculation was wrong. The fix is, unluckily, complicated – as far as I know, there are two common GISAXS geometries being used. This requires additional user choice here.
Here is the explanation; following pictures are from Lazzari, J. Appl. Cryst. (2002). 35, 406-421 and G. Renaud et al. / Surface Science Reports 64 (2009) 255–380):
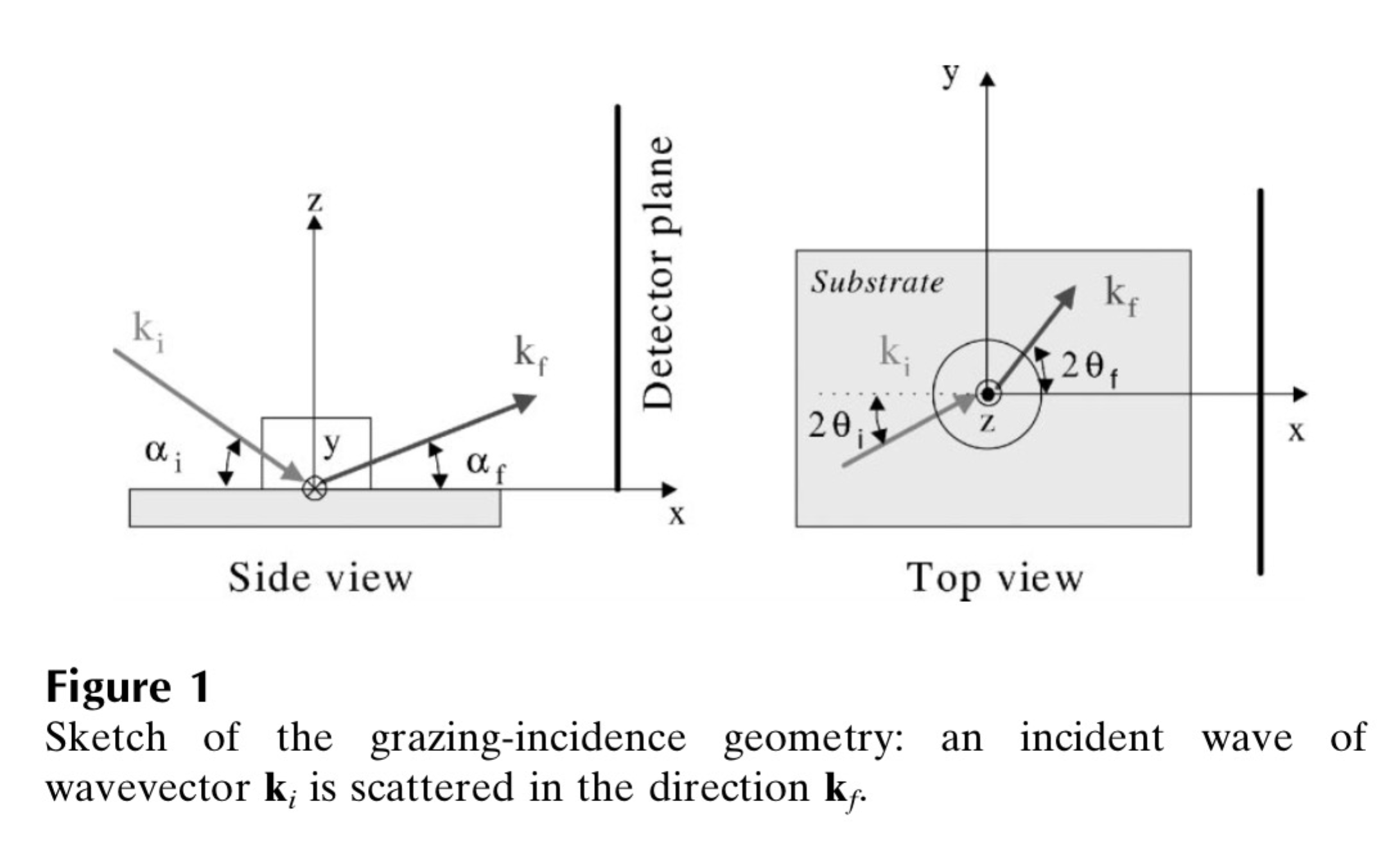
Here are the q components calculations based on this geometry. Note, Nika assumes Theta-I = 0.
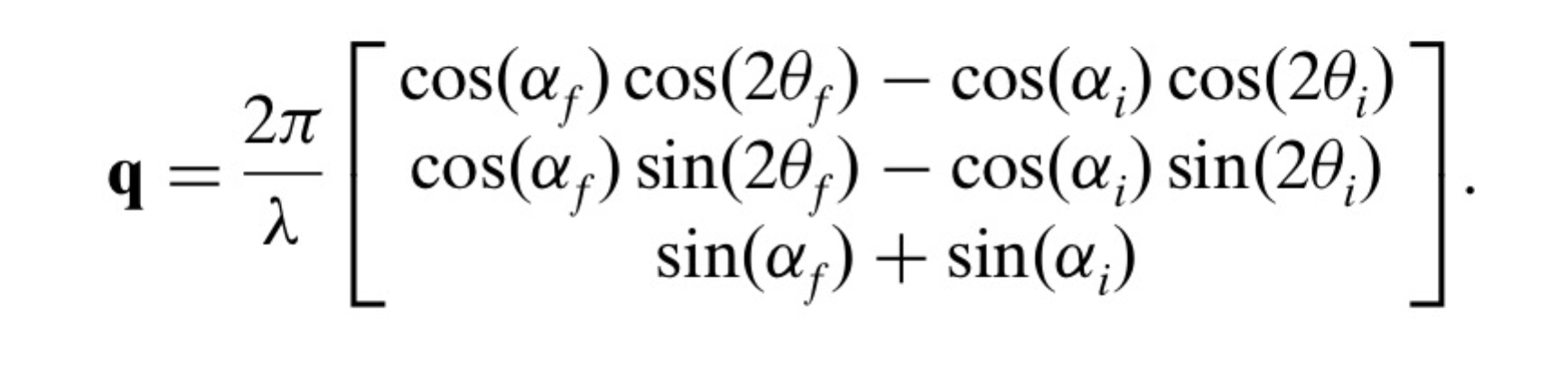
However, another geometry, which is also used, is slightly different:
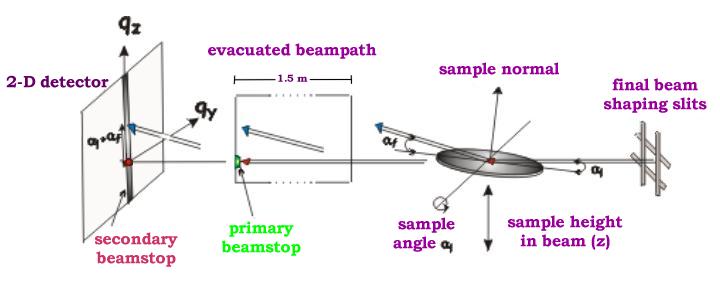
(Fig2. - http://www.physics.queensu.ca/~saxs/GISAXS.html)
Note the difference here is, that in the first image the sample is horizontal and beam is tilted, as it is commonly used for liquid surface scattering (“GEO_LSS”). For solid samples it may be more convenient to tilt the sample itself and rest of instrument stays fixed (“GEO_SOL”). In my rare encounters with GISAXS technique, this is what I have used.
These two geometries differ in the calculation of alfa-f needed for calculation of q in vertical direction. For GEO_SOL the detector is perpendicular to the original (incoming) beam direction and the alfa-f calculation does not require any more input from user as the calculation is simply the angle of the outgoing triangle – alfa-I as shown in Fig 2 here.
For the GEO_LSS as in Fig 1 the detector is perpendicular to the sample surface, and principally user should provide one more input parameter, as the triangles are not right angle any more. In this case users need to input another value – y position of the reflected beam.
Therefore if user selects GI geometry, from version 1.68 he/she should get new panel:
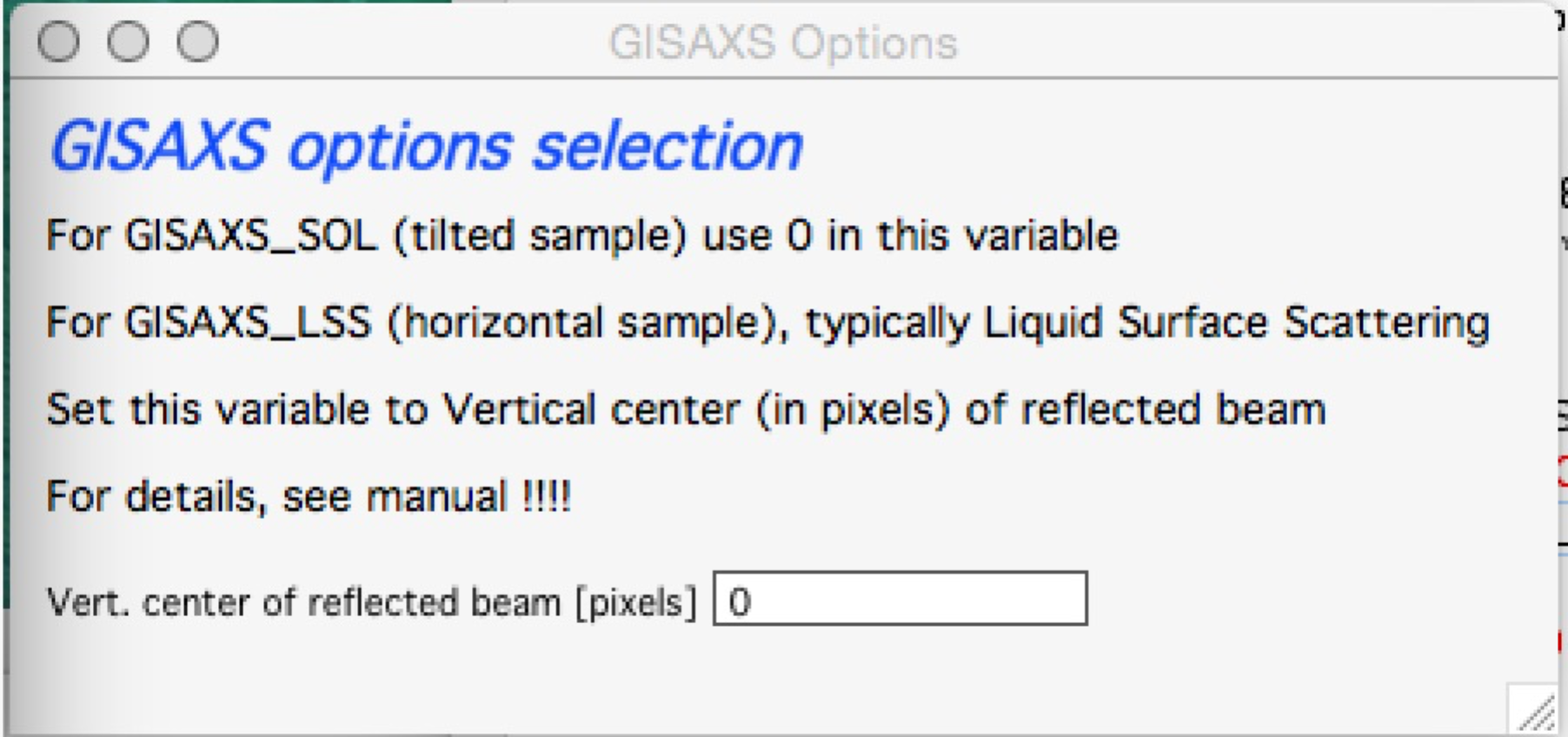
As instructed, for GISAXS_SOL where sample is tilted, just put (or leave) 0 in this field, close the panel and all is OK.
If you are using GISAXS_LSS geometry, you need to read (in pixels) position of the reflected beam and provide here the y coordinate of this beam. Close the panel and all should be set. Nika will use GISAXS_SOL calculation if this value is set to 0 (actually, if it is smaller than 1), and GISAXS_LSS if this value is larger than 0 (actually, >=1).
I do not have chance to test this, so if someone can test this and verify this all works, I would be really grateful.
And interestingly, there are instruments, which move their area detectors around much more, and orient them in much more complex way – and Nika has simply no chance to handle those systems. More complex instruments will require dedicated data reduction software.
The bug in this angle calculation was found by one of the users (Thank you!) in version 1.67 of Nika – the correction for alfa-I was missing.
** Ellipse profile**
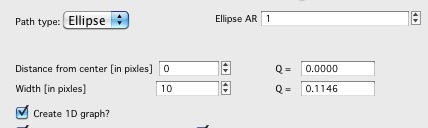
Note, that there is aspect ratio control here and the Distance from center here is horizontal distance (in qy) direction. When set to AR=1, the ellipse becomes circle.
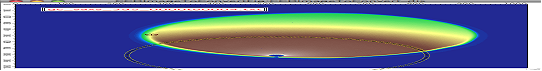

For AR>1, the ellipse is this way:
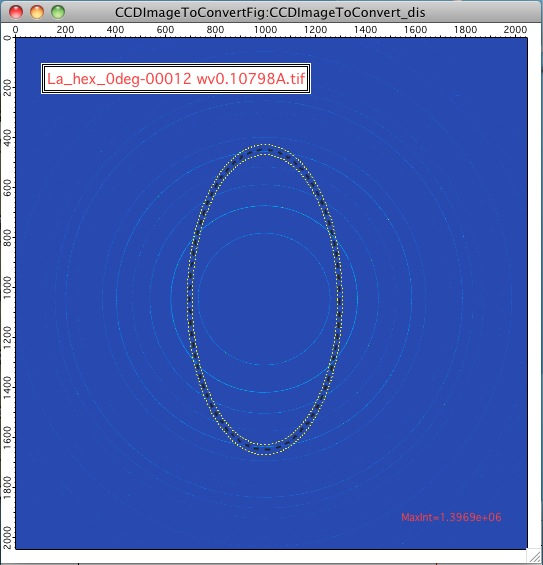
For AR<1, the ellipse is this way:
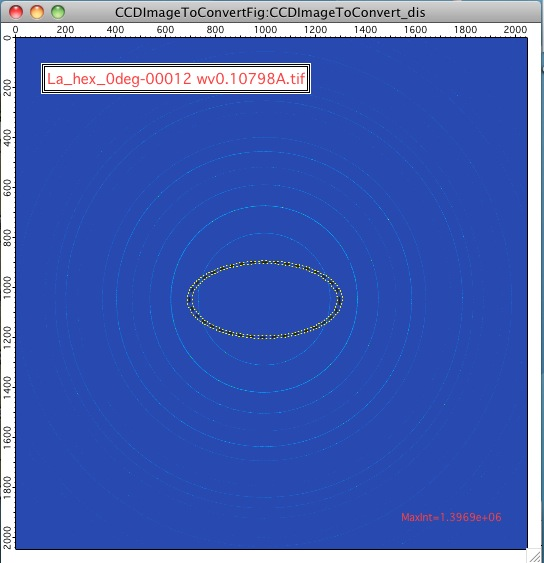
Note, that this tool has one major problem – it is practically impossible to display the data in any sensible way. Neither q, qz, or qy makes any sense here. In some way one needs to get angle of the intensity position. At this moment I do not produce such data within Nika. User can produce them by himself (the step is 0.25 degree, starting from 0 degrees azimuthal angle on the detector[note: I hope, I got turned around so many times, that this requires some data to test on]).
The other option is to use qy and qz to generate this angle. If anyone will ever use this tool, please, contact me and tell me, how you want to use it and I will modify the tool to suit needs of users.
*Finally : More shapes…. I can imagine broadening capabilities of this tool with other shapes. If you have such need, talk with me and I’ll add line profile shape for your needs. *
Controls for saving data are the same (really, these are the same controls, showing on second screen also) as in the Sectors tab:
Create 1D graph – if checked, 1d graph with output is created (if necessary) and data added. Note, the graph may be crowded very fast, since data are added, and added…
Store data in Igor experiment – keep data (as qrs triplets) in current Igor experiment.
Overwrite existing data if exist – if data with the same name exist, overwrite without asking. Otherwise, you will be asked.
Export data – export ASCII data
Select output path – select where data are to be placed.
Use input data name for output – automatically name 1D data (with sector information added as DataName_Angle_width) by input data name.
ASCII data name – if the above is not selected, this is place to put name for output file. Note, if there is nothing available for the code as sample name, it will ask for some…

Note, that the LineProf tool uses another “graph” window (“Line Profile Preview”) under the main image. This window contains some controls that are very useful.
The data are automatically updated as the parameters for the profile are changed. This gives user live update (but can take time, if it takes too much time for anyone, let me know and I’ll add controls to avoid the updates “live”).
User can display the data as function of q, q:sub:`y` or q:sub:`z` and on lin-lin, log-lin, lin-log and log-log scales. Note, that negative values cannot be displayed on log scale, so since q values for lower part of detector (below beam center) are defined as negative, you may not see them if you choose log scale. Also the q values look sometimes really weird, but generally they should be correct. If there are any issues with definitions of negative directions, let me know.
User can also save the data displayed in this window, which enables user to create multiple line profiles from existing image – this is manual method. NOTE that save parameters are taken from the setting of the controls for this purpose in the tab in the main panel (“Create 1D graph”, “Store data in Igor experiment”…). If you choose “Overwrite existing data” and do not change the name, you may get in troubles.
When data are being saved some cryptic description to indicate what profile was used and which q was used will be attached to the name used. More full description is attached to wave note.
For example for GI_Vertical line in my test case, this was the name:
gc_saxs_395__GI_VLp_0.0077
“gc_saxs_395_”…. Part of the name of used image
GI_VLp_…. GI_Vertical Line
0.0077 …. q:sub:`y` value at which the data were calculated.
Exported data are Int, error, Q, qx, qy, qz columns with header and column names
Saved data in Igor are
r_gc_saxs_395__GI_VLp_0.0077 intensity
q_gc_saxs_395__GI_VLp_0.0077 q
s_gc_saxs_395__GI_VLp_0.0077 error
qy_gc_saxs_395__GI_VLp_0.0077 qy
qz_gc_saxs_395__GI_VLp_0.0077 qz
qx_gc_saxs_395__GI_VLp_0.0077 qx (generated ONLY if GI… profile is used)
Note: next release of Irena package will have capabilities to use not only qrs data , but also qxrs, qyrs, and qzrs data.
Save tab¶
This tab is intended to control all data saving in Nika.
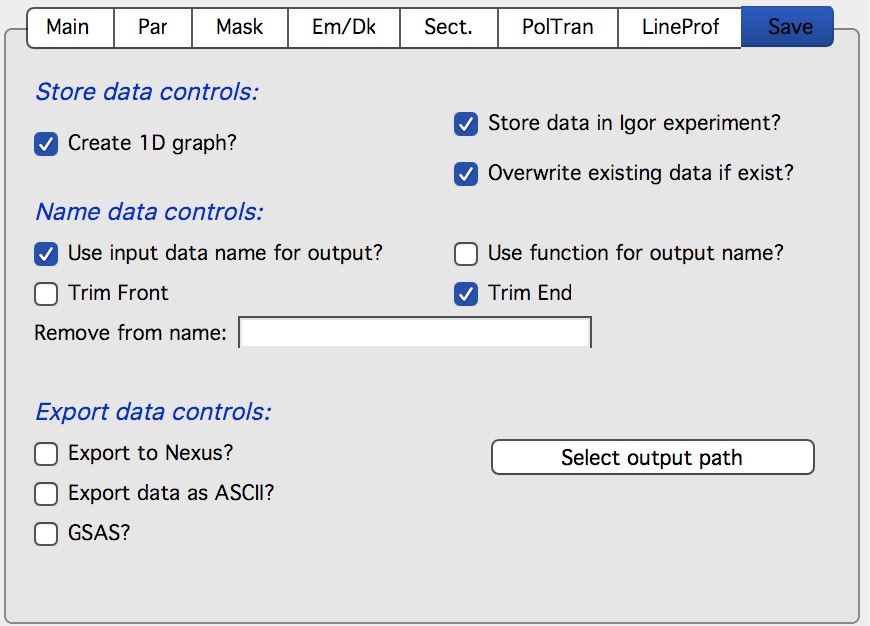
Store data controls:
This controls how data re stored in Igor. Note, that if you want to Create 1D graph, Store data in Igor must be checked also and the code will do so. Overwrite existing data if exists will overwrite data in Igor if the same name data already exist. If you process same data set multiple times for testing, check it. If you process many files, uncheck it as this will prevent accidental overwriting of the data in case their names end up to be same.
Name data controls:
This controls how data are named when imported. Here is where user can select how the names are created using either imported image name with optionally some trimming, user can write Igor Function which will provide the necessary name etc.
Export data controls:
This controls if the data will be automatically exported as they are processed. This is quite useful if other programs, such as sasView, are going to be used for data analysis. Options are
Export to ASCII This allows users to ASCII data (4 columns, Q, Intensity, Uncertainty and Qresolution) into text files.
GSAS Should export GSAS compatible ASCII data.
Export to Nexus This allows users to export NXcanSAS 1D data to NEXUS. As of version 1.80 this should work for sasView. This required some modifications which I did not expect as sasView cannot load standard NXcanSAS file, the file has to be quite specific.
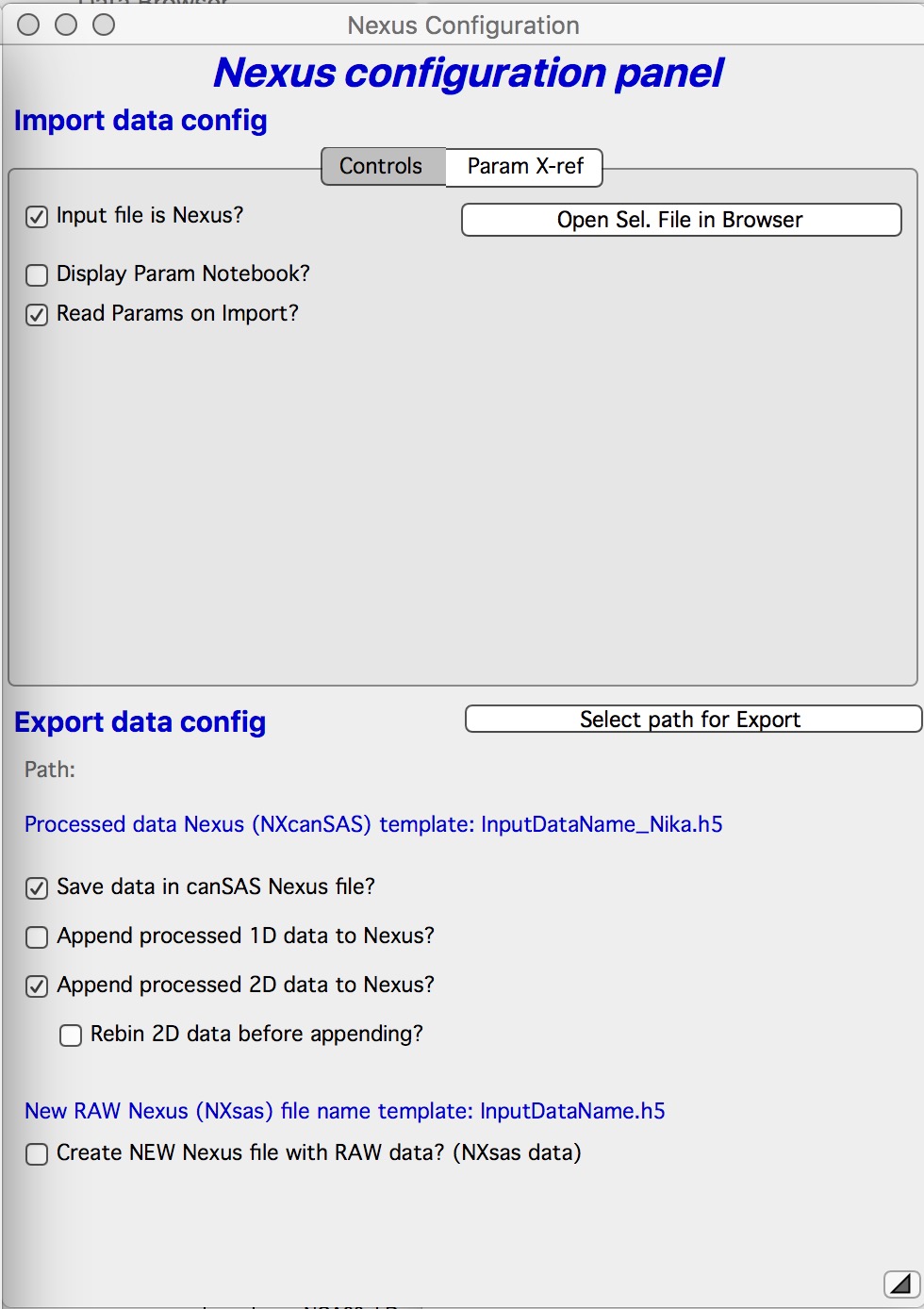
Checking the checkbox Export Nexus brings up dialog for Nexus Export and import. In this case the important part is Bottom part. User needs to setup export path (folder on drive) and select what will be exported. SUGGESTION use ONLY the top option “Save data in canSAS Nexus File”. This will create individual Nexus file for each data set. Second option “Append processed 1D data to Nexus” will append multiple data sets in single file. When I tried this, file with ~40 images hang sasView badly and I needed to kill it.
“Append processed 2D data to Nexus” will append the calibrated 2D data in the file. It is not obvious if there is any program which can accept these data, so, even though there is standard on this, it makes little to no sense to do.
Create NEW Nexus (NXsas) file with RAW data” this is interesting option, it allows you to take tiff file or any other source file Nika can read and make it into RAW data nexus file. Note, this is RAW data file with metadata Nika knows. This should be possible then to use for further data reduction. Not sure how useful and meaningful this is.
Bottom controls¶

These controls have following functions:
“Display only checkbox” Nika will average all selected files, which are selected in the list box, and display them as one image. The program will just load and display the images, including some processing (dezinging), if selected. but no calibration or other processing is done. This is really for preview of how the image looks like.
Note, if more than 1 image is selected, the images are first AVERAGED – that is intensities for each pixel as summed together and then divided by number of images.
“Process sel. files individually” Nika will load one image at a time from the files selected in the list box and processeach individually according to selection in the tabbed area. For each input file you get all output data (whatever you selected above).
“Avergae all selected and process” Nika will average all selected files in the list box and process them - together as one input data - according to selection in the tabbed area. You get ONE output data (whatever you selected above) for all together. Typically used when multiple image of same condition are collected to improve statistics.
Note, if more than 1 image is selected, the images are first AVERAGED – that is intensities for each pixel as summed together and then divided by number of images.
“Average N of selected and process” Nika will average Sequentially (in order) N selected files in the list box and process them - together as one input data - according to selection in the tabbed area. You get ONE output data (whatever you selected above) for each N images. Typically used when multiple image of same condition are collected to improve statistics.
This opens further controls:
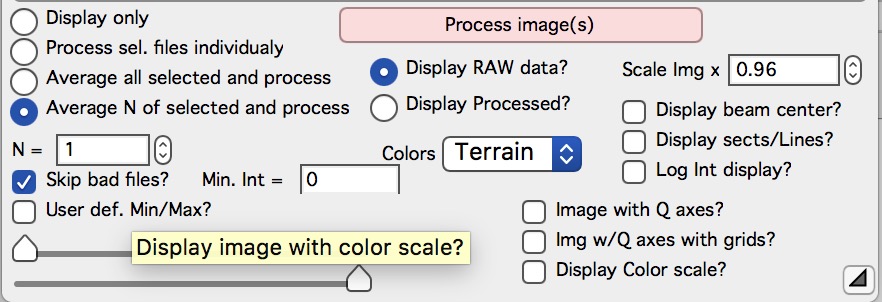
“N =” This controls how many images Nika will avergae over.
“Skip Bad files” Enables to skip automatically processing of files, which have too low intensity (SetVariable control with limiting value appears when selected). Used to skip files which were accidentally NOT exposed in case of failing shutters or other issues.
“Min int =” This defines “bad image”. Typically bad image has much lower intensity than good image (shutter did not open, instrument failed) and so one one set minimum intensity in image needed to consider such image a good one. If bad image is found, it is skipped. Note, that even bad images are counted in the “N” value.
“Display RAW data” will display in the image right of the panel the UNCORRECTED data file as loaded in. Values for the pixles are raw counts from the detector.
“Display Processed” will display in the image right of the panel the fully CORRECTED and CALIBRATED data. The values for the pixles should be directly absolute intensity in this case. This choice is not available, if image was loaded through using “Ave & Display sel. Files(s)”. In this case no processing of the image was done. Use button “Convert sel. Files 1 at time” or the other buttons…. Just remember, that only the last image is available for display.
“Colors” Choice of color scales. These are now remembered on a given computer, the the last one should be reused next time. Default is Terrain.
“Scale Img x” User can select how large the image shouLd be displayed on the screen. If input image is too large, set smaller so it fits on the screen (this should eb done automatically anyway), if it is small, scale up to have it cover larger fraction of the screen.
“Display beam center” will add circles in the image showing where beam center is set
“Display sectors/Lines” will add lines showing sectors or lines, which are selected for data analysis (if any)
“Log Int display” will switch displayed image into log (intensity) or linear (Intensity).
“image with Q axes” Appends Qx/Qy (or Qz/Qy) axes to displayed image. Note, when unchecked, it has to recreate the image, since these Q axes cannot be removed any other way.
“image w/ Q axes with grid” Appends Qx/Qy (or Qz/Qy) axes to displayed image – with grid lines. Note, when unchecked, it has to recreate the image, since these Q axes cannot be removed any other way.
“Display Color Scale?” Appends color scale to image.
“User def. Min/Max?” Opens controls to set manually max and min intensity to display in the image. Does not change when new image is loaded.
“Sliders” Slide to set min and max intensity displayed in the image. Resets when new image is loaded.
Polarization correction¶
Two types are available.
Unpolarized radiation
This is generally accepted formula.
Linearly polarized radiation
This is polarization correction for linearly polarized radiation, such as produced by double-crystal monochromators on synchrotrons.
There are two polarization orientations, sigma (linear part) and pi. Most synchrotrons will be linearly sigma polarized, with sigma fraction may be 0.99 or so. Depending on the way the detector is read, the sigma polarization plane may be horizontal or vertical. The panel enables setting the sigma polarization plane orientation.
The final formula is:
where fs is fraction of sigma polarization, 2q is 2 theta angle, and a is azimuthal angle from the plane of polarization plane.
Implementation
All of the Polarization corrections (from version 1.42) in Nika are applied by scaling the 2D data by the formulas above after all of the corrections (including background and dark current subtraction).
In the following panel which shows after selecting “Polarization correction” on the main panel:
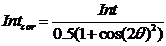
After selecting Polarized radiation you need to make further choice…
If the Sigma Polarization Plane is 0 degrees, then the detector orientation is such, that the polarization plane is horizontal in the Nika image of the detector. Note that horizontal is Nika’s definition of 0 degrees on the detector.
This has nothing to do with the orientation of polarization in real World, this is an orientation between the polarization plane and the way detector is read. In this case the correction looks like this:
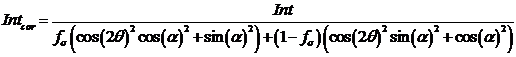
with largest correction (increase of intensity) where the color is blue.
For case, when polarization plane is vertical in Igor image (perpendicular to Nika’s definition of 0 degrees on detector) , the correction looks like this:

with maximum correction (blue color).
Uncertainties (“Errors”)¶
Uncertainty estimate in 2D data reduction is sore point and I have not yet found correct solution for it. As far as I know there is really no good way to get meaningful estimates.
To complicate the matter is, that prior version 1.43 (1.42 and before) there is bug in the uncertainty (error) calculation, which results in overestimate of the values. My intention was to provide standard deviation of the values averaged into the pixel, but simply, I made typo, which resulted in somehow higher values.
Therefore for version 1.43 I provide now three different methods for uncertainity calculations, Standard deviation is default. For compatibility purposes user can choose old (incorrect) version and also standard error of mean – SEM - (standard deviation / sqrt(number of points)).
Please note, that the line profile calculations provide ONLY standard deviation or SEM, since they never used the old method (they use Igor internal method for standard deviation). They default to standard deviation if old method is selected.
The Uncertainty method can be changed in the “Configuration panel” available from menu.

Q-resolution calculations¶
From Nika version 1.69 the code can estimate q-resolution of the data. This is highly approximate calculation, which can be probably, similar to Uncertainties calculations considered voodoo calculations. I have reviewed some manuscripts which deal with this , such as Barker, J. Appl. Cryst (1995) 28, 105-114. I have looked in some of the codes and realized, that while this is challenge to do for a specific instrument (USAXS code handles this as correctly as anyone probably ever will need), for generic tool this will be challenge. And to some degree, for X-ray instruments this is mostly (not always!) OK as the resolutions are kind of higher than what neutron system need to deal with.
Here is description of what Nika does to calculate q resolution for each point.
Wavelength resolution is ignored. For regular monochromatic instruments this is reliably ignorable value. For pink beam, well, if you need it I can add it in the future, but I am not sure if anyone needs it (and this would require yet another GUI control value few people would ever use). So if you need it, let me know and we will deal with it then.
Effect of q-binning. When Nika calculates intensity, it calculates q value for center of each pixel and then generates q binning (linear or logarithmic) – this means, each q-bin has qmin and qmax. All pixels with qcenter between qmin and qmax are counted for each bin. Nika provides this q-width (distance between qmin and qmax) as q resolution given by nature of averaging.
Effect of pixel size. Note, that above the q is placed into the bin based on center q value. Of course, this means, that some pixels with center near qmin or qmax contain intensity from q values belonging to other q bins due to finite pixel size. This is q resolution due to pixel size.
Effect of beam size. Now one needs to realize, that beam has finite size and often is really large. Therefore each pixel will see range of q values (angles) from different places on the beam spot. At the end, this is very similar to pixel size smearing but with beam size values. This is q resolution given by beam size.
Effect of detector pixel bleeding. This is caused by detectors not being able to separate the intensity in one pixel from the next pixel. This is highly detector technology dependent and Nika simply ignores it. Luckily, newer generations of detectors (Pilatus) are pretty good in this.

Note, that adding the Beam size q-resolution required adding of controls for the beam size into the main GUI. If beam size is left as 0, the only thing affected is the q-resolution calculation. This is beam size ON DETECTOR! not on the sample. If there is focusing, that can cause differences.
OK, so in the table above (and that is not exhaustive table) are some of the sources of the q resolution we need to account for. Nika convolutes together Effect of q-binning, effect of pixel size and effect of beam size. It ignores others.
There are bit more details in how the calculations are handled and in case of real interest, read the code (the function is NI1A_CalculateQresolution in NI1_ConvProc.ipf). It gets bit messy in the way these things get expressed:
For “small” q-resolution values caused mainly by pixel size and beam size – and where the q-binning is smallish (or at least comparable) component, the correct is expressing q-resolution as FWHM (full width of half maximum) of assumed Gaussian sensitivity of the q bin across of range of q values. This is what most software assumes. This is what you get always at small qs in Nika.
For “large” q widths generated at high-q by log-q binning in Nika (and in USAXS using flyscans etc.) the correct representation is more as rectangular slit smearing effect (similar to slit smeared USAXS instrument itself). This is what you get if you use Nika with log-q binning at higher qs.
Irena Modeling II has been recently updated to handle this type of q-smearing. It is bit mess for number of options
Summary:
Accounting for q-resolution can be helpful for scattering with sharp features (monodispered systems etc…). It may be critical for fitting such systems as I was unable to fit some of these systems without accounting for q-resolution. Keep that in mind when fitting is not going well.
It can also be very useful to look at to decide what is the real q minimum value of any instrument. I have seen cases when device is quoted to have qmin – 0.0006 A-1 but the q resolution at that pixel is about 0.002 A-1, which really makes that pixel useless for practical purposes. I think this is more common than we dare to accept…
Recently updated Modeling II tool in Irena can handle different types of q-smearing.