Important Information¶
The code uses for all size related parameters Angstroems (10-10 m) or for Q vector (A-1). In the case of scattering contrast, number distribution and any other volume contents centimeters (10-2 m).
Output of the size is usually in particle diameters, but Modeling II is using radii as default (and diameters optionally), but read the graphs, the output may not be always the same. Output graph legend or panel text should be always correct.
Loading the macros¶
Start Igor Pro
In menu “Macros” select “Load Irena SAS macros”.
Irena code is loaded and compiled. This can take a while. New menu “SAS” appears. This is where all the Irena macros are controlled from.
Unload the macros¶
When you want to pass Igor experiment to someone without Irena installed, it is good idea to remove the Irena from current experiment. Removing the macros itself is achieved by selecting “Remove Irena package”, the very last item in the last “SAS” submenu. This will unload macros and put back in the “Macros” menu command to load Irena macros in the future.
Data naming conventions¶
Data in Irena need to be stored within Igor experiment as “waves” which is Igor term for “array” of data. Typical SAXS or WAXS data will have x-axis (q vector) and y-axis (intensity). Optionally we can have other information - intensity uncertainty, q-resolution etc. These are “input” data for Irena, typically either generated by Nika or Indra data reduction packages or imported from other data reduction packages using one of the Data Import tools.
Not only we have “input” data, but most Irena tools will generate some “output” data - e.g., Size distribution, Unified fit model intensity vs Q, etc. These are “Irena results”.
Irena uses strictly “Folder = sample” philosophy. It means, that within Igor experiment (to see it graphically, open Data Browser - in Igor menu “Data”, select “Data Browser”) is located folder which name represents your sample name. Only one data set per folder. Most Irena operations will pick the data from this folder and if user decides to save the results from any modeling, return proper Irena result back to this folder. If more than one set of data is in a folder, it is impossible to figure out which data the result belongs to.
Data manipulations tools, which create a new (e.g., merged) data set will create a new folder.
Data naming conventions - QRS¶
This is data naming convention for data stored within Igor experiment, typically in folder root:SAS: in folders with folder name reflecting the sample name.
The wave names
X-axis data
q_NameOfSample q vector in A-1
t_NameOfSample 2 theta, if output with respect to 2 theta in degrees
d_NameOfSample d for output with respect to d in A
y axis data
r_NameOfSample intensity (if calibrated in whatever units – thickness is converted to cm, so it should be cm-1)
error
s_NameOfSample error for intensity
other
w_NameOfSample width of each bin of Q, d, or 2 theta.
Data naming conventions - QIS¶
This is data naming convention for data stored within Igor experiment, typically in folder root:SAS: in folders with folder name reflecting the sample name. This is NIST modification to QRS system and Irena treats this “transparently” as QRS system.
The wave names
X-axis data
NameOfSample_q q vector in A-1
y axis data
NameOfSample_i intensity (if calibrated in whatever units – thickness is converted to cm, so it should be cm-1)
error
NameOfSample_s error for intensity
Data naming conventions - USAXS¶
This is data naming convention for data stored within Igor experiment, required to be in folder root:USAXS: in folders with folder name reflecting the sample name. Note, that slit smearing is reflected in the names and Irena will understand the difference. Irena switches on slit smearing based on the names.
The wave names
X-axis data
SMR_Qvec (slit smeared) - DSM_Qvec (pihole collimated) q vector in A-1
y axis data
SMR_Int (slit smeared) - DSM_Int (pihole collimated) intensity (if calibrated in whatever units – thickness is converted to cm, so it should be cm-1)
error
SMR_error (slit smeared) - DSM_error (pihole collimated) error for intensity
other
SMR_dQ (slit smeared) - DSM_dQ (pihole collimated) width of each bin of Q, d, or 2 theta.
Slit smeared (USAXS) data¶
Fitting slit smeared data is major Irena advantage. It is nearly ALWAYS better to fit slit smeared data than to desmear the data and then fit them. However, some Irena tools have limitations when fitting smeared data: due to need to smear data and the way it is handled by Irena routine, the modeled range of data (the high-q selected for modeling and fitting) have to be larger than slit length. Typical slit length of the USAXS instrument is 0.02-0.03 A-1, so the high-q range needed to be at least 0.08 A-1. This means, that it is not possible to select for modeling data from small-qs to only 0.02 A-1 ONLY.
This is fixed now for Modeling, Unified Fit, and Size distribution by adding additional Q points (up to 100 new points) to extend the data to 10* slit length and after the model is calculated and slit smeared, the data are truncated back to original user range. This is done automatically, behind user back – but note, that it can cost cpu and therefore increase time per calculation of the model. Therefore, it may be worthwhile to simply select high enough Qmax for fitting anyway.
Per point smeared data by Q resolution¶
New in version 2.58 and only for Modeling at this time, check the manual. This is a beast of issues but can be important!
Kill all Irena panels and graphs¶
This menu item allows closing all Irena related windows – panels and graphs – to be closed at once. Very convenient…
Irena manual¶
In most cases button Help present on most panels should open Irena manual in web browser on web site:
http://saxs-igorcodedocs.readthedocs.io/en/latest/index.html
You can also download from there version of the manual in pdf, epub, and html as zip file. These can be used for off-lien reading of manual.
Irena youtube channel¶
Movies are available here :
https://www.youtube.com/channel/UCDTzjGr3mAbRi3O4DJG7xHA/about
Open Form factor description¶
This should open pdf file with form factors description – including simplified code and graphs. These are form factors in the “central bank” of the Irena, available for use in packages, which use them.
Check for updates¶
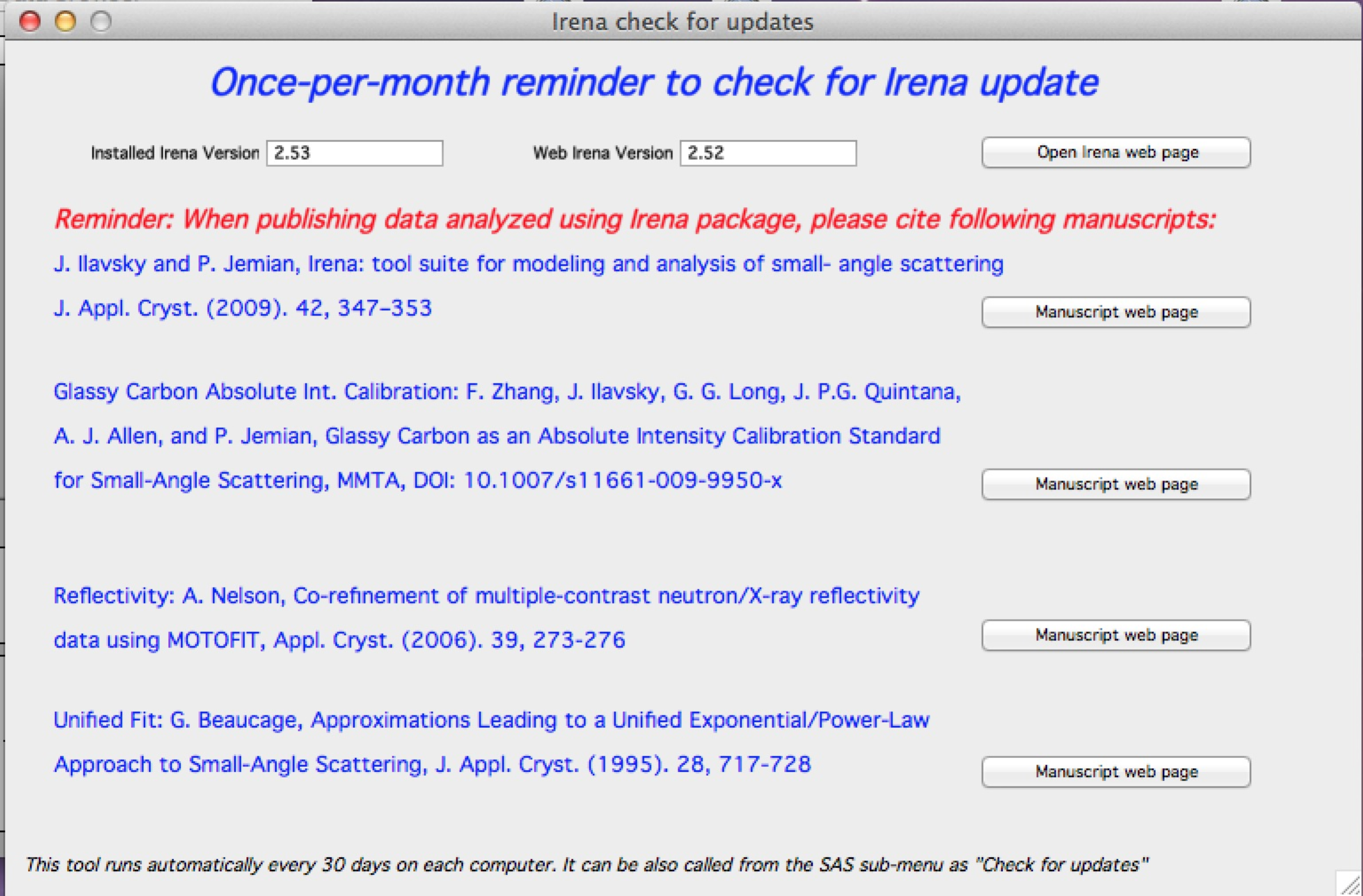
From version 2.52 I have added once-per month check for updates, which on ANY computer runs every 30 days. It checks installed versions of the packages and web available versions. It also reminds you about need to cite manuscripts related to the Irena and tools implemented in Irena. Please, cite those manuscripts as necessary.
You can get this panel opened any time from SAS>Help, About,…> Check for updates
The buttons open appropriate web pages in your web browser.
GUI controls and common controls¶
Manual, Manuscript, Mailing list, About…
From the Last menu Item you can get “About” panel stating current version and Igor versions, which it has been tested on.

Download and open Manual, request manuscript, sign up for mailing list and do few other operations you may find useful. Including “offloading” Irena package from the experiment, so it does not slow down the operations when you want to do something else. Or when you want to send file to someone who may not have Irena installed, remove Irena package so he/she does not get errors on load when Igor tries to load Irena unsuccessfully.
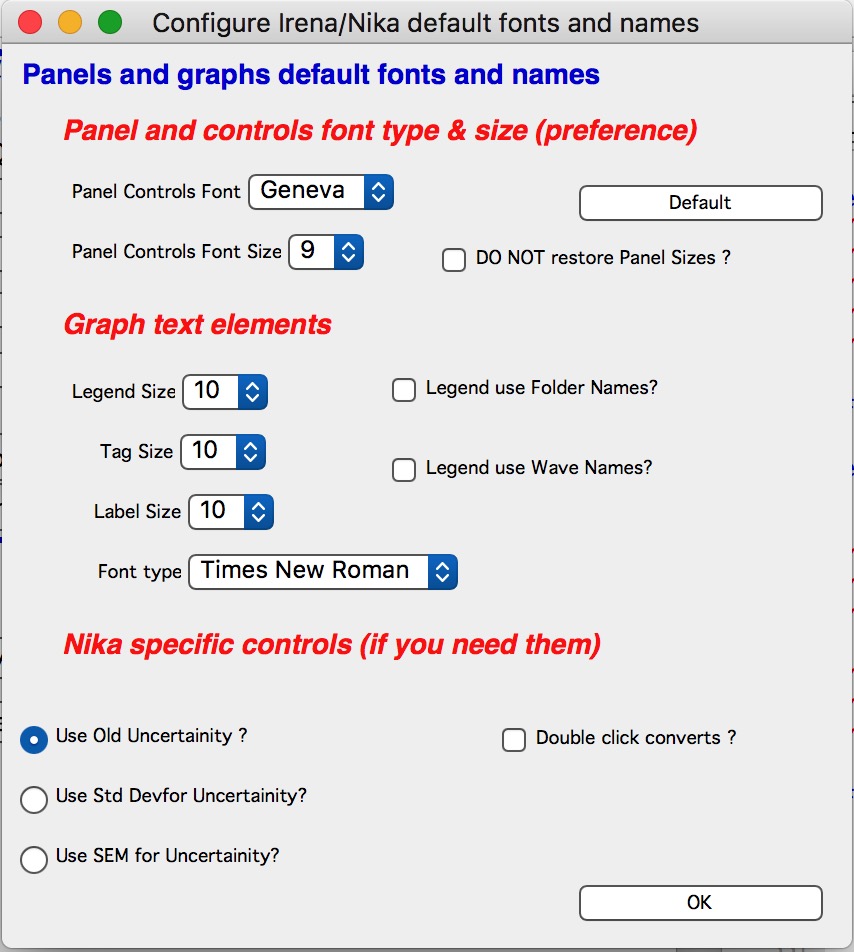
Configure Figure default fonts and names
“Configure Figure default fonts and names” in the SAS menu will create panel with some controls common for all tools, like font type & size and how legend names are handled. NOTE: Panel controls are applied immediately to all existing panels, graph controls are applied ONLY to the newly created graphs (and only those which were upgraded to this behavior).
DO NOT restore Panel Sizes
Do NOT restore Panel Sizes” controls if panels are restored to the last used size and position when other they are recreated (they were closed and a tool is opened again) or when some existing experiment is reopened. Keep in mind, that if this checkbox is not selected, every time you scale up/down a panel, its position and size is recorded. When being recreated, the panel will move and scale back to its size. NOTE: Position and size is recorded ONLY when panel size is changed, not when it is just moved. If you want to overwrite hti sbehavior, hold down any modifier key (alt, cmd/ctrl/shift…).
Panels font and font sizes
These controls enable user to customize font used on control panels therefore this enables customization for a given platform. This is necessary as more and more control is provided on each platform to user and therefore default fonts and font sizes may not be appropriate any more for the panels I design. These settings are actually saved on a given machine as well as the experiment. This has some interesting features, so please, read carefully:
When these controls are run (and user is forced to run them if the Irena is loaded and preferences are not found), they save preferences in special folder Igor maintains for users. At the same time, the settings are applied to the current experiment.
When this experiment is opened on another computer, the preferences from that computer are not reloaded, so the experiment will use preferences from the original computer. When the “Configure image GUI and Graph defaults” is run, it will reload the computer defaults and apply them to the given experiment. Then user can change the fonts and font sizes as they wish. The new settings are saved on the computer – and within the experiment.
Note, that Panel font and font size are platform specific, so same experiment may present differently looking panels on Mac and PC. Also, from version 2.62 this panel is common for Irena and Nika packages, so not everything you see in Irena applies.
Note, not all controls actually follow these settings, I have been changing some buttons to specific font and font size and those are not affected by these settings.
If there are any issues with the behavior, please, let me know and I’ll see if I can make it more logical.
Note the difference in ConFigure GUI and Graph defaults panels when different fonts are used. You can mess up the panels really well by wrong choices!
Defaults button returns the panel font choices to platform specific default state (Mac: Geneva size 9 and PC Tahoma size 12). Note, that there is no guarantee that these were your choices before. But these should be reasonable choices for most setups.
Graph controls
I am slowly adding in various parts of the whole package calls to these commonly stored values. This allows user to conFigure fonts for various screen sizes. This seems necessary to allow use of Mac/Win platforms with vastly different screen sizes and resolutions.
Not all packages follow these controls yet, if you see issues in package of your choice, let me know and I will try to address them ASAP. Time is limited resource.
Data selection¶
Data selection part of the panels is served by common package (mostly) and has more or less similar behavior – with modifications appropriate for each package. The purpose of these controls is to provide as much help to user to select appropriate data as possible. This is not easy task… Sometimes even it is not clear what the right help is.
There are few checkboxes for data types, up to 4 popups with Data Folder, Wave with X, Y and error data. If Model input is appropriate, Qmin, Qmax, number of points and log/lin binning inputs are displayed.
How the control works:
Type of data:
Indra 2 data data from Indra package (DSM_Int, etc.). Assumes data are in root:USAXS folder (or any subfolder) only.
QRS data data with q_name, r_name (intensity) and optionally s_name (error). Alternatively, to help users using NIST SANS data analysis package the option recognizes also “qis” system (“name_q”, “name_i”, “name_s”) and presents the data with this naming system as well. NOTE: Irena now carries forward, if present, also w_name wave, assumed to contain dq values. This is created by Nika or Data import packages and can be used for per-pixel smearing in Modeling II package. While it does not show in any GUI, if present, it is handled correctly.
Model No data, tool will create q data using user input and intensity/error data will be set to 0. Then passed intot he tool so one can model with no measured data present. Available ONLY when appropriate.
Irena results should know results from Irena package (all different types). When appropriate will be available. Note, that in any folder may be number of different results available.
User type currently not used, but allows definition of any other naming structure to be used in the future. Note this can be named differently at any time and can provide access to any doublet or triplet of wave types, if it can be defined.
No type of data selected In this case the tool will present choice of all folder in the experiment and for data waves all of the wave in the particular folder. This method will work always, but may be quite challenging to use.
Basic control logic
When particular type of data is selected, the tool should go and find all of the folders containing at least one of the type of data.
Indra 2 data at least one of M_DSM_Int (M_DSM_Qvec, M_DSM_Error), DSM_Int, M_SMR_Int, SMR_Int triplets.
QRS data triplet of waves starting with q, r, s with the rest of name the same. Note, this is the most cpu challenging data type, so it will take the longest.
Irena results any of the results from Irena package. If any is missing, let me know, please…
Model no input data, input data will be created.
User not used at this time. Can be used in the future for any data types, which can be defined.
Nothing all folders, all waves available.
These folders are presented in the “Data folder” for user selection. When user selects the folder, rest of Wave popups will be populated by first valid set, which is in the order prescribed by internal logic.
If other data set is needed, select different data in the “Wave with X axis data” popup. This will attempt to fill the next ones with appropriate data. This may not be unique, so the first match will be filled in.
Then if still necessary, fill in the other two popups.
Note, that it is possible, that depending on tool you can select only two data waves (X and Y), some tools may require also error wave.
Folder/Wave name masking :
Starting with Irena 2.53 I have enabled use of “weird” characters in names - (){}%#^$?|&@ can now be used as part of the name… This modified option to mask Folder name and/or Wave name with string to make smaller selection in the popups. There are two new fields now – and yes, it is possible the new string fields get hidden below controls for Folder and Q wave selection. There is not enough space, select “—“ in that popup to get to these new controls.
Since version 2.53 these controls allow user to only string to match the names to select folder/waves to be displayed. Prior version enabled use of Regex, but since now control characters for Regex are part of the name and hence possibly part of the match string, it is now impossible to use Regex and one has to use simple string. DO NOT add * if you want to match part of the name, simply using string “test” will match any name which has anywhere in it test as string.
Little useful trick: Regular expression which means “not matching string xyz” is ^((?!xyz).)*$ - yes, it is weird, but works. Replace xyz by string of characters contained in data which you do not want to have displayed and they will disappear from the list.
Here is how to use it:

This is how the default state looks – empty field for “Fldr” and “Wvs”. If there is empty string, all folders and waves of that specific type will be presented.
See here, we have 4 samples measured and we have now 4 folders available.
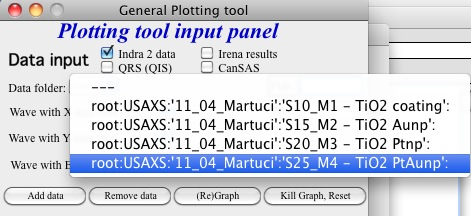
Here is setting when I want to match Aunp string to be in each of the names:
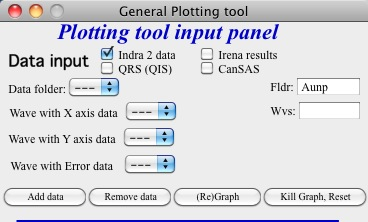
and here is what is presented as result of the above choice:
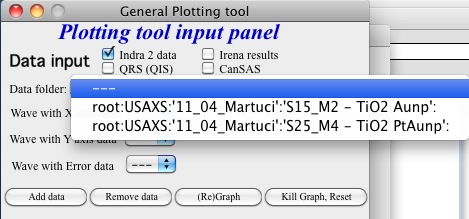
Little help:
Typical use is to show only data with specific match string, to display only selections, which contain “abcd” in the name just put the abcd letters in the field. No * are necessary.
If you want to use two strings which a name must contain, use this : String1.*String2. Keep in mind that String1 must occur before String2 in the name to be matched. And yes, between them is “.*” without any spaces.
Match strings are tool-specific, so each tool has its own specific set of match strings.
MultiData selection¶
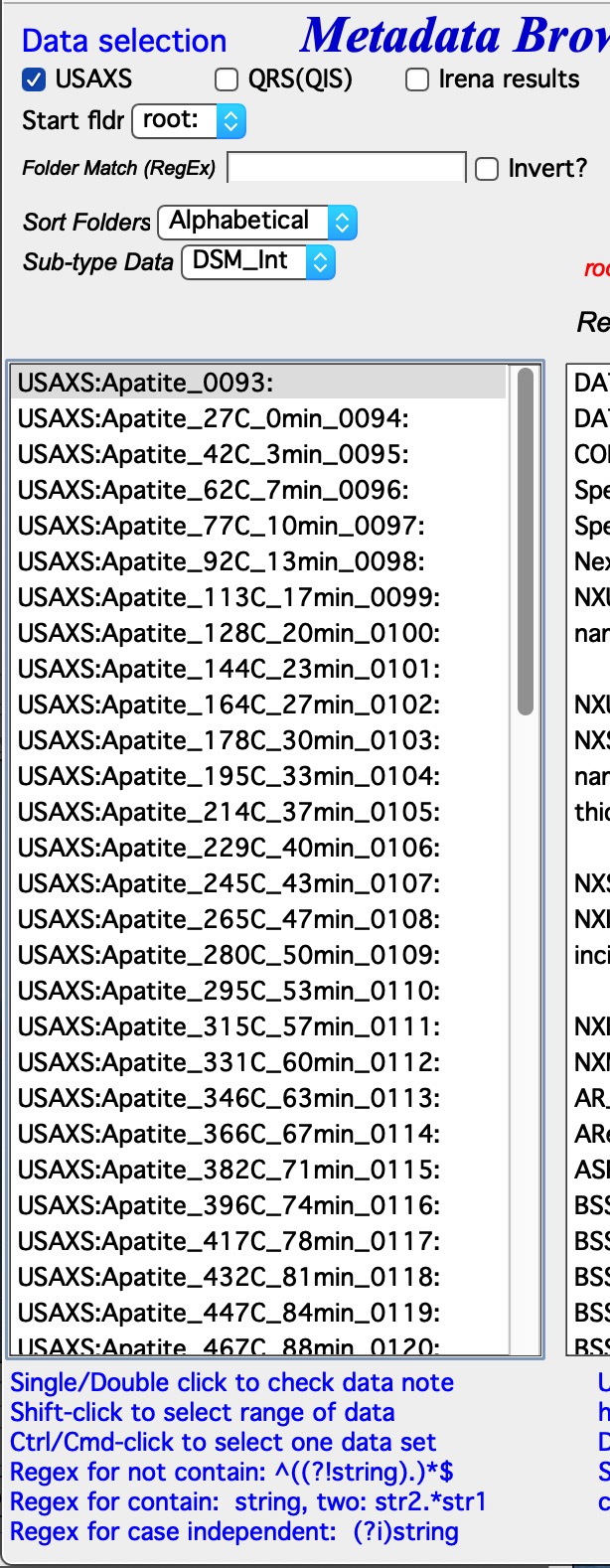
Data type Irena recognizes few data types.
USAXS data type = this is naming system for data generated by APS USAXS instrument. Ignore, unless you have data from this instrument. If you have our data, you should know enough to use this.
QRS data type = this is the default data naming system for SAXS/WAXS data in Irena and Nika. For details see here QRS data type.
Irena results = any fit and modeling results generated by Irena. Most tools will save some type of data - size distribution, fits, pddf, diffraction peaks,… All of these data types can be seen as “Irena results”
Any = if all checkboxes are unchecked, user can define Regular expressing, which will tell irena which wave is x, y, and optionally error. Keep in mind, that the first wave matching the regular expression will be picked. This may require some testing or help from me, if you want to use it.
Start Fldr. Here you can select at which location in data tree code will start looking for the data. Pick suitable place, for example root:SAXS may be a good start. Picking suitable start where to look for data makes the code run faster.
Folder Match (RegEx) this allows users to look for only some of the folders. A short summary on regular expressions is at the bottom of the page, below the Listbox with folder. Google it, understanding regular expressions will be very helpful.
Invert? this checkbox inverts the Regular expression meaning. So if you insert in the “Folder Match” field string 00034, only data which have in name 00034 will show. If you check this checkbox, selection is inverted and all files which do NOT contain this string in the name will show.
Sort Folders This sorts the folders using one of many methods implemented. As result, this will group folders in order which may be helpful for processing. For example, some tools create list of results in the order the samples were processed. Having proper order helps plotting results after the analysis properly.
Interesting and complicated are results In this case there are many choices and user needs to be very careful and will have to pick lots of choices.
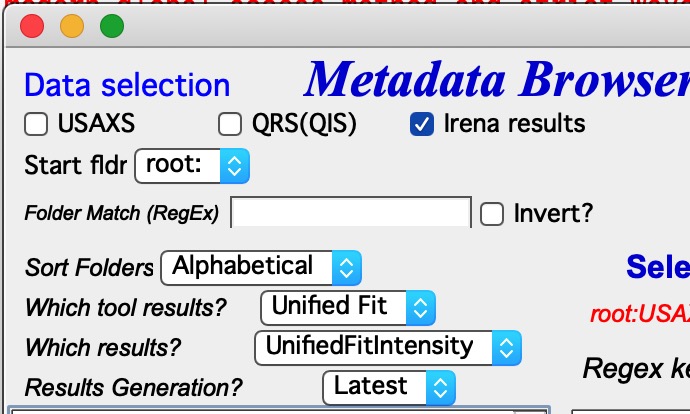
Which tool results is selection of Irena tools, in this example it is Unified fit. Code will show only folders which contain some Unified fit results. But there are many results which are possible.
Which results will offer choices of type of results, for example UnifiedFitIntensity. Now, keep in mind that if you save Unified fit results multiple times, you will get what is called “generations” so you need to pick also which generation to use.
Results generation options are Latest, and from _0 to _10.
And even more complicated are no defined data selection In this case user needs to provide regular expression, which provides way to get uniquely wave in folder. In the image you can see regular expressions to match QRS naming system. But, this is very flexible and user can principally design suitable regular expressions for whatever is needed.

HOW TO USE Pick Data selection checkboxes properly. Pick a good Starting folder. If needed, use Folder Match (RegEx) to see only data folders you want to see. Sort the data based on whatever is appropriate in the Sort Folders options. Learn some regular expressions, handy cheatsheet is below the Listbox… Regular expressions, at least basics, help a lot!
Using Irena on small displays¶
Irena generates a lot of windows, panels, graphs, notebooks… It really needs large display, 1024x768 is simply too small for useful work. Current version of Irena requires at least 1100 x 900 pixels display - but this is much more complicated on Widowns with the high resolution displays…
To aid users I have now added function which calculates what the area available for content is (in Igor pixel units). On start my code now checks and if available area is smaller than preset values (1100 x 900) the code provides warning in a dialog and instructions in History area. The code will still work, but some tools may refuse to run since the panels would not fit on the screen. On Windows users can change the display resolution and/or reduce the Display screen settings (“dpi”). On Macs users can increase the display resolution. To recheck the size after changing the settings, use command “Check Igor display size” from the menu USAXS, SAS2D, or SAS>”Help, About, Manuals, Remove Irena”. For more details if you have problems seeing panel content, please, see GUI Controls Missing in Common Issues.
GET LARGE ENOUGH DISPLAY. THEY ARE CHEAP NOW…
It is possible to move the content (not all, but most) up/down on panels, when needed with the arrows in top right corner:
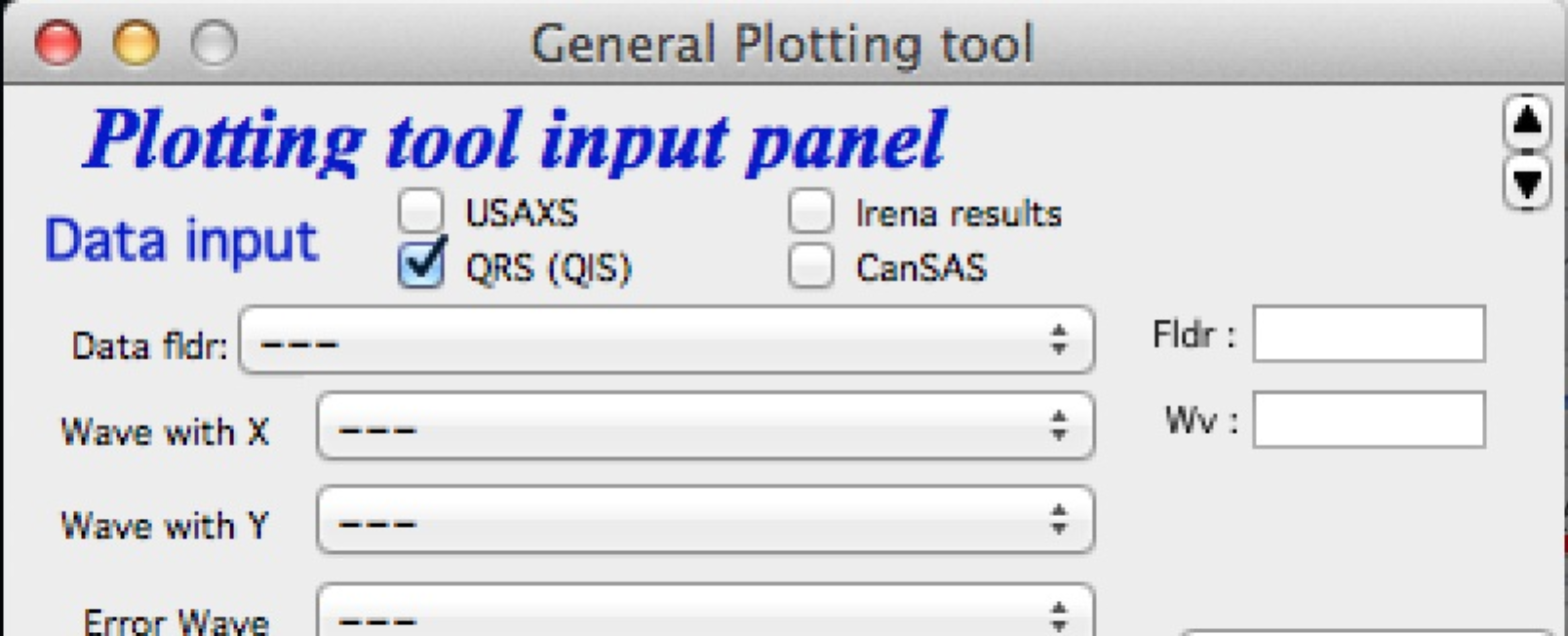
The two arrows at the top right corner of most panels - like here on plotting tool panel - move the content of the panel up/down, so if your screen is too small vertically (usual problem), you can move the controls in the screen itself. However, this is a chllenge in Igor 7 and does not work too well.
So here is the same area, but content was now moved bit higher, so one can reach to the bottom controls:
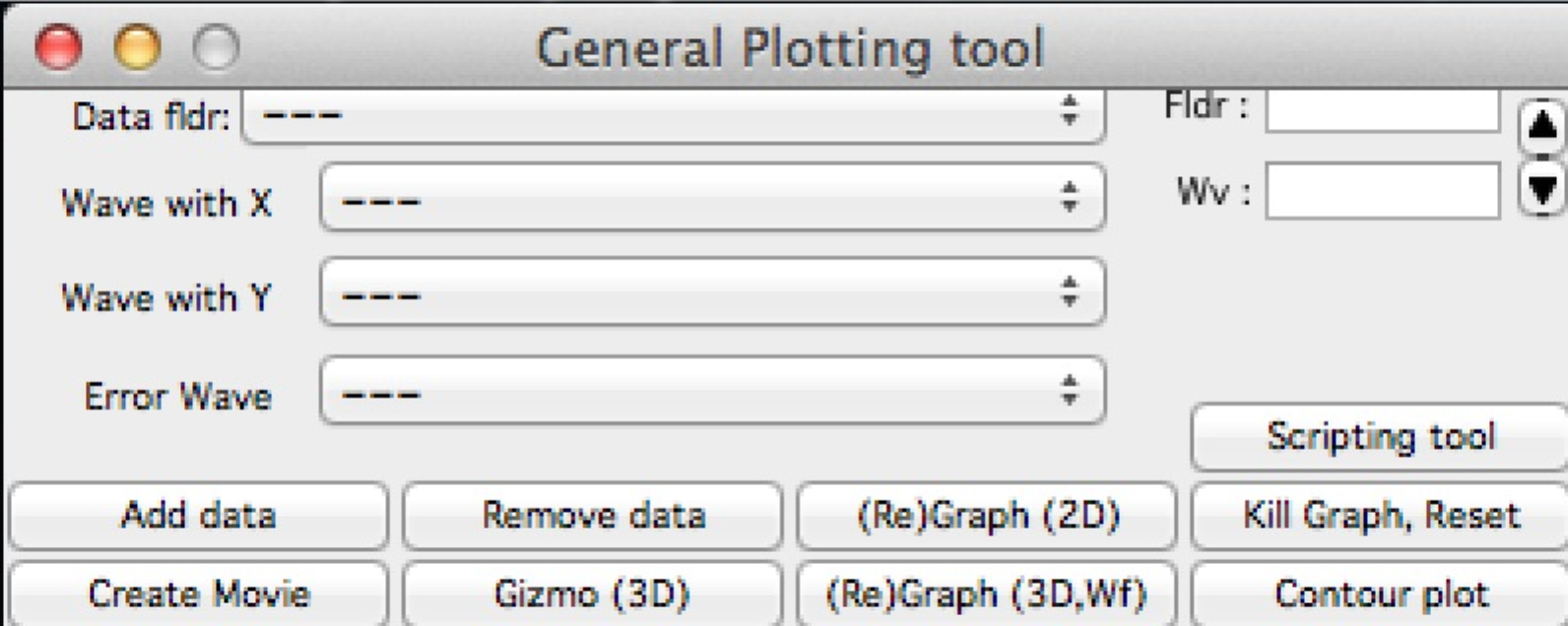
If you have a large display, you can zoom panels by dragging lower right corner - note mark:

You can scale panels up or down, but they will not scale to smaller size than original size.
NOTE: the setting of size and position is now persistent - it is recorded as preference on the current computer. Therefore, if you scale panel up and then close the panel, next time you recreate this panel, it will be rescaled for you to the same size. Panel size and position will also be restored when an experiment is re-opened. This can cause panels to move around with respect to the positions in which they were stored when the Igor experiment was saved last time. If you do not like this, change the setting as indicated below.
However, for usability in case you changed the display size in the mean time, the panel will be also imited in size to 50% width fo the current display AND 90% height of the current display. If you want to reset the panel to its default size, hold down shift/alt or cmd/ctrl key while creating the panel again. The size will be reset.
There is override to thsi behavior in the Configure defaults (see above) panel, where you can choose not to have panel size and position restored.
Using Irena on high resolution displays¶
Use of XOP¶
Igor Pro enables use of external C-code to speed up some high cpu intensive operations. Note, that these binary pieces of code and bit-specific, so there is specific version for Igor 32bit and specific for Igor 64bit versions. They need to be properly located in Igor folder structure. Currently various optional xop program are available:
Two by Andrew Nelson http://motofit.sourceforge.net/wiki/index.php/Main_Page – one for calculation of reflectivity (abeles.xop) and one for genetic optimization (GenCurvefit.xop). Both are compulsory (for functionality of Reflectivity and Genetic optimization) and need to be placed in “Igor extensions” folder. Both speed up the calculations by factor of up to 40 compared to now removed Igor code. They need to be kept updated, so please, update with every new Irena update as they do not have version numbers.
XML loader (also by Andrew Nelson) necessary to load XML (CanSAS) file formats. You can download this general use XML xop from : http://www.igorexchange.com/project/XMLutils
Version 2.53 added first form factor (Parallelepiped) which is available ONLY xop library maintained by NIST reactor. Version 2.54 and higher can take advantage of speed improvements for some other form factor also (cylinder, spheroid). NIST colleagues (Steven Kline namely) were nice enough to provide me with updated versions of their xops and I suggest you use the ones available with my package.
Genetic optimization¶
Genetic optimization method is form of fitting from SAS data. It has been developed for optimization of reflectivity data but is very useful for cases where least square fitting may not find global minimum. It has been programmed for Igor by Andrew Nelson, who is also author of internal code for reflectivity tool.
Note that this code uses some version of Monte Carlo method. Therefore limits are _very_ important. When Genetic optimization method is used user will be presented with dialog to check the limits. For this method is really important that the calculations do not fail for any combination of parameters and that the range of probed parameters is sensible.